Focus and Scope
Biodiversity Data Journal (BDJ) is a community peer-reviewed, open access, comprehensive online platform, designed to accelerate publishing, dissemination and sharing of biodiversity-related data of any kind. All structural elements of the articles – text, morphological descriptions, occurrences, data tables etc. – will be treated and stored as data, in accordance with the Data Publishing Policies and Guidelines of Pensoft Publishers.
The journal will publish papers in biodiversity science containing taxonomic, floristic/faunistic, morphological, genomic, phylogenetic, ecological or environmental data on any taxon of any geological era from any part of the world with no lower or upper limit to manuscript size. For example:
- single taxon treatments and nomenclatural acts (e.g., new taxa, new taxon names, new synonyms, changes in taxonomic status, re-descriptions, etc.);
- data papers describing biodiversity-related databases, including ecological and environmental data;
- sampling reports, local observations or occasional inventories, if these contain novel data;
- local or regional checklists and inventories;
- habitat-based checklists and inventories;
- ecological and biological observations of species and communities;
- any kind of identification keys, from conventional dichotomous to multi-access interactive online keys;
- descriptions of biodiversity-related software tools.
For more information, you may look at the Editorial Beyond dead trees: integrating the scientific process in the Biodiversity Data Journal and press release The Biodiversity Data Journal: Readable by humans and machines.
Globally Unique Innovations
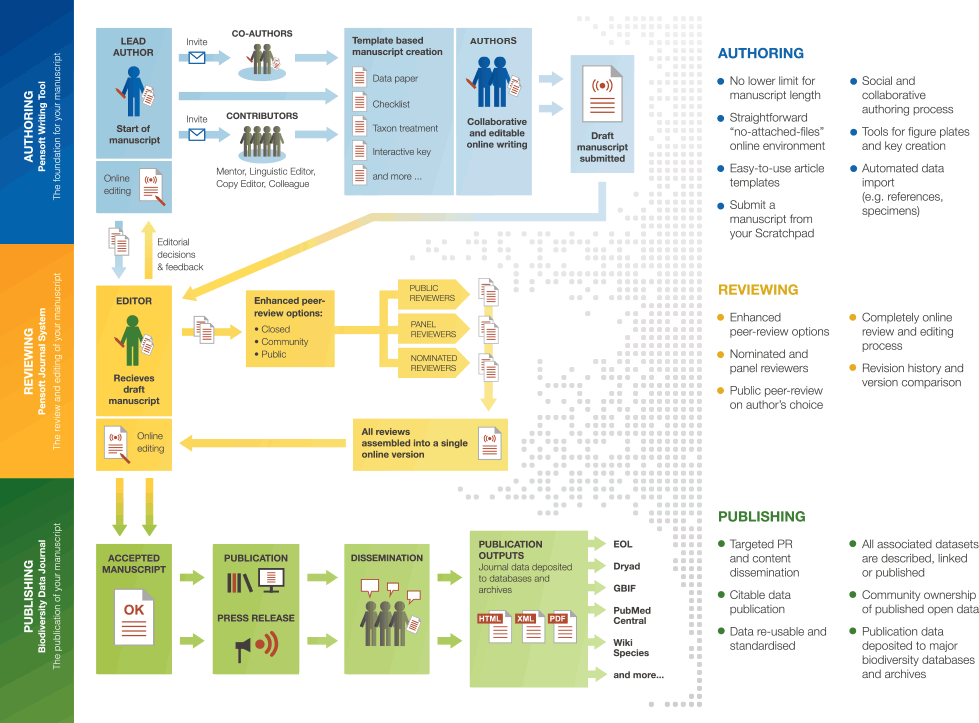
The Biodiversity Data Journal (BDJ) and the associated ARPHA Writing Tool (formerly Pensoft Writing Tool), launched within the FP7 project ViBRANT, created several, globally unique, innovations:
- The first work flow ever to support the full life cycle of a manuscript, from writing through submission, community peer review, publication and dissemination within a single online collaborative platform.
- The online, collaborative, article-authoring platform ARPHA Writing Tool (AWT) provides a large set of pre-defined, but flexible, Biological Codes and Darwin Core compliant, article templates.
- Authors may work collaboratively on a manuscript and invite external contributors, such as mentors, potential reviewers, linguistic and copy editors, colleagues, who may monitor and comment on the text before submission. These comments can be submitted along with the manuscript for editor’s consideration.
- Import/export conversion of data files into text and vice versa, from text to data, such as checklists, catalogues and occurrence data in Darwin Core format, simply at the click of a button.
- Automated import of data-structured manuscripts generated in various platforms (Scratchpads, GBIF Integrated Publishing Toolkit (IPT), authors’ databases).
- A novel community-based pre-publication peer-review and possibilities to comment after publication (post-publication peer-review). Authors may also opt for an entirely public peer-review process. Reviewers may opt to be anonymous or to disclose their names.
For more information, you may look at the Editorial Beyond dead trees: integrating the scientific process in the Biodiversity Data Journal and press release The Biodiversity Data Journal: Readable by humans and machines.
Copyright Notice
License and Copyright Agreement
In submitting the manuscript to any of Pensoft’s journals, authors certify that:
- They are authorized by their co-authors to enter into these arrangements.
- The work described has not been published before (except in the form of an abstract or as part of a published lecture, review or thesis); it is not under consideration for publication elsewhere; its publication has been approved by all author(s) and responsible authorities – tacitly or explicitly – of the institutes where the work has been carried out.
- They secure the right to reproduce any material that has already been published or where copyright is owned by someone else.
- They agree to the following license and copyright agreement:
Copyright
- Copyright on any article is retained by the author(s) or the author's employer. Regarding copyright transfers please see below.
- Authors grant Pensoft Publishers a license to publish the article and identify itself as the original publisher.
- Authors grant any third party the right to use the article freely as long as its original authors and citation details are identified.
- The article and published supplementary material are distributed under the Creative Commons Attribution License (CC BY 4.0):
Creative Commons Attribution License (CC BY 4.0)
Anyone is free:
to Share — to copy, distribute and transmit the work
to Remix — to adapt the work
Under the following conditions:
Attribution. The original authors must be given credit.
- For any reuse or distribution, it must be made clear to others what the license terms of this work are.
- Any of these conditions can be waived if the copyright holders give permission.
- Nothing in this license impairs or restricts the author's moral rights.
The full legal code of this license.
Copyright Transfers
Any usage rights are regulated through the Creative Commons License. Since Pensoft Publishers is using the Creative Commons Attribution License (CC BY 4.0), anyone (the author, their institution/company, the publisher and the public) is free to copy, distribute, transmit and adapt the work as long as the original author is credited (see above). Therefore, specific usage rights cannot be reserved by the author or their institution/company and the publisher cannot include a statement "all rights reserved" in any published paper.
Website design and publishing framework: Copyright © Pensoft Publishers.
CLOCKSS system has permission to ingest, preserve, and serve this Archival Unit.
Authorship/Contributorship
Some journals are integrated with Contributor Role Taxonomy (CRediT), in order to recognise individual author input within a publication, thereby ensuring professional and ethical conduct, while avoiding authorship disputes, gift / ghost authorship and similar pressing issues in academic publishing.
During manuscript submission, the submitting author is strongly recommended to specify a contributor role for each of co-author, i.e. Conceptualization, Methodology, Software, Validation, Formal analysis, Investigation, Resources, Data Curation, Writing - Original draft, Writing - Review and Editing, Visualization, Supervision, Project administration, Funding Acquisition (see more). For the journals that are not integrated with CRediT, the submitting author is encouraged to specify the roles as a free text. Once published the article will include the contributor role for all authors in the article metadata.
Desk Rejection
During the pre-review evaluation, Editors-in-Chief or Subject editors check the manuscript for compliance with the journal's guidelines, focus, and scope. At this point, they may reject a manuscript prior to sending it out for peer review, specifying the reasons. The most common ones are non-conformity with the journal's focus, scope and policies and/or low scientific or linguistic quality. In such cases, authors are encouraged to considerably improve their manuscript and resubmit it for a review. We encourage authors whose manuscripts have been desk rejected due to being out of the scope of this journal to consider another potentially suitable title from the Pensoft portfolio.
In case the manuscript is suitable for the journal but has to be corrected technically or linguistically, it will be returned to the authors for improvement. The authors will not need to re-submit the manuscript but only to upload the corrected file(s) to their existing submission.
Peer Review
This journal uses a single-blind peer review process. This means that the names of reviewers are hidden from the authors (the author does not know the identity of the reviewer, but the reviewer knows the identity of the author). Notwithstanding that, the reviewers are encouraged to disclose their identities, if they wish to do so. Each article is reviewed by at least two independent experts, with a final decision on acceptance being made by the Subject Editor / Editor-in-Chief. Front-matter articles, such as editorials, correspondence, biographies, and similar articles, can be published after editorial evaluation only.
Please consider the Editor and Reviewer Guidelines in the About webpage of this journal for more details and stepwise instructions on the editorial and peer review process.
Preprints
This journal allows posting preprints of the manuscripts submitted for peer-review. Authors are strongly encouraged to use the ARPHA Preprints server for that as an option available during the submission process, which will save a double effort in manuscript submission and allows the preprint to be directly linked to the published article and vice versa.
Manuscripts that contain nomenclatural acts in the sense of the biological Codes will not be posted as preprints even when the authors opt for that, to avoid possible confusion in the priority of names and validity of publication.
Indexing and Archiving
The articles published in the journal are indexed by a high number of industry leading indexers and repositories. The journal content is archived in CLOCKSS, Zenodo, Portico and other international archives. The full list of indexes and archives are shown on the journal homepage.
The authors are allowed to publish preprints of their manuscripts on ARPHA Preprints or other preprint servers. The deposition and distribution of preprints and final article versions is highly encouraged.
Advertising
Publication Ethics and Malpractice Statement
General
The publishing ethics and malpractice policies follow the Principles of Transparency and Best Practice in Scholarly Publishing (joint statement by COPE, DOAJ, WAME, and OASPA), the NISO Recommended Practices for the Presentation and Identification of E-Journals (PIE-J), and, where relevant, the Recommendations for the Conduct, Reporting, Editing, and Publication of Scholarly Work in Medical Journals from ICMJE.
Privacy statement
The personal information used on this website is to be used exclusively for the stated purposes of each particular journal. It will not be made available for any other purpose or to any other party.
Open access
Pensoft and ARPHA-hosted journals adhere strictly to gold open access to accelerate the barrier-free dissemination of scientific knowledge. All published articles are made freely available to read, download, and distribute immediately upon publication, given that the original source and authors are cited (Creative Commons Attribution License (CC BY 4.0)).
Open data publishing and sharing
Pensoft and ARPHA encourage open data publication and sharing, in accordance with Panton’s Principles and FAIR Data Principles. For the domain of biodiversity-related publications Pensoft has specially developed extended Data Publishing Policies and Guidelines for Biodiversity Data. Specific data publishing guidelines are available on the journal website.
Data can be published in various ways, such as preservation in data repositories linked to the respective article or as data files or packages supplementary to the article. Datasets should be deposited in an appropriate, trusted repository and the associated identifier (URL or DOI) of the dataset(s) must be included in the data resources section of the article. Reference(s) to datasets should also be included in the reference list of the article with DOIs (where available). Where no discipline-specific data repository exists authors should deposit their datasets in a general repository such as, for example Zenodo or others.
Submission, peer review and editorial process
The peer review and editorial processes are facilitated through an online editorial system and a set of email notifications. Pensoft journals’ websites display stepwise description of the editorial process and list all necessary instructions and links. These links are also included in the respective email notification.
General: Publication and authorship
- All submitted papers are subject to a rigorous peer review process by at least two international reviewers who are experts in the scientific field of the particular paper.
- The factors that are taken into account in review are relevance, soundness, significance, originality, readability and language.
- A declaration of potential Conflicts of Interest is a mandatory step in the submission process. The declaration becomes part of the article metadata and is displayed in both the PDF and HTML versions of the article.
- The journals allow several rounds of review of a manuscript. The ultimate responsibility for editorial decisions lies with the respective Subject Editor and, in some cases, with the Editor-in-Chief. All appeals should be directed to the Editor-in-Chief, who may decide to seek advice among the Subject Editors and Reviewers.
- The possible decisions include: (1) Accept, (2) Minor revisions, (3) Major revisions, (4) Reject, but re-submission encouraged and (5) Reject.
- If Authors are encouraged to revise and re-submit a submission, there is no guarantee that the revised submission will be accepted.
- The paper acceptance is constrained by such legal requirements as shall then be in force regarding libel, copyright infringement and plagiarism.
- No research can be included in more than one publication.
- Editors-in-Chief, managing editors and their deputies are strongly recommended to limit the amount of papers co-authored by them. As a rule of thumb, research papers (co-)authored by Editors-in-Chief, managing editors and their deputies must not exceed 20% of the publications a year, with a clear task to drop this proportion below 15%. By adopting this practice, the journal is taking extra precaution to avoid endogeny and conflicts of interest, while ensuring the editorial decision-making process remains transparent and fair.
- Editors-in-Chief, managing editors and handling editors are not allowed to handle manuscripts co-authored by them.
Responsibility of Authors
- Authors are required to agree that their paper will be published in open access under the Creative Commons Attribution License (CC BY 4.0) license.
- Authors must certify that their manuscripts are their original work.
- Authors must certify that the manuscript has not previously been published elsewhere.
- Authors must certify that the manuscript is not currently being considered for publication elsewhere.
- Authors should submit the manuscript in linguistically and grammatically correct English and formatted in accordance with the journal’s Author Guidelines.
- Authors must participate in the peer review process.
- Authors are obliged to provide retractions or corrections of mistakes.
- All Authors mentioned are expected to have significantly contributed to the research.
- Authors must notify the Editors of any conflicts of interest.
- Authors must identify all sources used in the creation of their manuscript.
- Authors must report any errors they discover in their published paper to the Editors.
- Authors should acknowledge all significant funders of the research pertaining to their article and list all relevant competing interests.
- Other sources of support for publications should also be clearly identified in the manuscript, usually in an acknowledgement (e.g. funding for the article processing charge; language editing or editorial assistance).
- The corresponding author should provide the declaration of any conflicts of interest on behalf of all authors. Conflicts of interest may be associated with employment, sources of funding, personal financial interests, membership of relevant organisations or others.
- Manuscripts in revision have to be revised and resubmitted within a reasonable time span. The authors are aware that manuscripts not revised within 100 days after the revision decision will be rejected and have, if desired by the authors, to be submitted afresh.
Responsibility of Reviewers
- The manuscripts will be reviewed by two or three experts in order to reach first decision as soon as possible. Reviewers do not need to sign their reports but are welcome to do so. They are also asked to declare any conflicts of interests.
- Reviewers are not expected to provide a thorough linguistic editing or copyediting of a manuscript, but to focus on its scientific quality, as well as for the overall style, which should correspond to the good practices in clear and concise academic writing. If Reviewers recognize that a manuscript requires linguistic edits, they should inform both Authors and Editor in the report.
- Reviewers are asked to check whether the manuscript is scientifically sound and coherent, how interesting it is and whether the quality of the writing is acceptable.
- In cases of strong disagreement between the reviews or between the Authors and Reviewers, the Editors can judge these according to their expertise or seek advice from a member of the journal's Editorial Board.
- Reviewers are also asked to indicate which articles they consider to be especially interesting or significant. These articles may be given greater prominence and greater external publicity, including press releases addressed to science journalists and mass media.
- During a second review round, the Reviewer may be asked by the Subject Editor to evaluate the revised version of the manuscript with regards to Reviewer’s recommendations submitted during the first review round.
- Reviewers are asked to be polite and constructive in their reports. Reports that may be insulting or uninformative will be rescinded.
- Reviewers are asked to start their report with a very brief summary of the reviewed paper. This will help the Editors and Authors see whether the reviewer correctly understood the paper or whether a report might be based on misunderstanding.
- Further, Reviewers are asked to comment on originality, structure and previous research: (1) Is the paper sufficiently novel and does it contribute to a better understanding of the topic under scrutiny? Is the work rather confirmatory and repetitive? (2) Is the introduction clear and concise? Does it place the work into the context that is necessary for a reader to comprehend the aims, hypotheses tested, experimental design or methods? Are Material and Methods clearly described and sufficiently explained? Are reasons given when choosing one method over another one from a set of comparable methods? Are the results clearly but concisely described? Do they relate to the topic outlined in the introduction? Do they follow a logical sequence? Does the discussion place the paper in scientific context and go a step beyond the current scientific knowledge on the basis of the results? Are competing hypotheses or theories reasonably related to each other and properly discussed? Do conclusions seem reasonable? Is previous research adequately incorporated into the paper? Are references complete, necessary and accurate? Is there any sign that substantial parts of the paper were copies of other works?
- Reviewers should not review manuscripts in which they have conflicts of interest resulting from competitive, collaborative, or other relationships or connections with any of the authors, companies, or institutions connected to the papers.
- Reviewers should keep all information regarding papers confidential and treat them as privileged information.
- Reviewers should express their views clearly with supporting arguments.
- Reviewers should identify relevant published work that has not been cited by the authors.
- Reviewers should also call to the Editors’ attention any substantial similarity or overlap between the manuscript under consideration and any other published paper of which they have personal knowledge.
Responsibility of Editors
- Editors in Pensoft’s journals carry the main responsibility for the scientific quality of the published papers and base their decisions solely on the papers' importance, originality, clarity and relevance to publication's scope.
- The Subject Editor takes the final decision on a manuscript’s acceptance or rejection and his/her name is listed as "Academic Editor" in the header of each article.
- The Subject Editors are not expected to provide a thorough linguistic editing or copyediting of a manuscript, but to focus on its scientific quality, as well as the overall style, which should correspond to the good practices in clear and concise academic writing.
- Editors are expected to spot small errors in orthography or stylistic during the editing process and correct them.
- Editors should always consider the needs of the Authors and the Readers when attempting to improve the publication.
- Editors should guarantee the quality of the papers and the integrity of the academic record.
- Editors should preserve the anonymity of Reviewers, unless the latter decide to disclose their identities.
- Editors should ensure that all research material they publish conforms to internationally accepted ethical guidelines.
- Editors should act if they suspect misconduct and make all reasonable attempts to obtain a resolution to the problem.
- Editors should not reject papers based on suspicions, they should have proof of misconduct.
- Editors should not allow any conflicts of interest between Authors, Reviewers and Board Members.
- Editors are allowed to publish a limited proportion of papers per year co-authored by them, after considering some extra precautions to avoid an impression of impropriety, endogeny, conflicts of interest and ensure that the editorial decision-making process is transparent and fair.
- Editors-in-Chief, managing editors and handling editors are not allowed to handle manuscripts co-authored by them.
Neutrality to geopolitical disputes
General
The strict policy of Pensoft and its journals is to stay neutral to any political or territorial dispute. Authors should depoliticize their studies by avoiding provoking remarks, disputable geopolitical statements and controversial map designations; disputable territories should be referred to as well-recognised and non-controversial geographical areas. Тhe journal reserves the right to mark such areas at least as disputable at or after publication, to publish editor's notes, or to reject/retract the paper.
Authors' affiliations
Pensoft does not take decisions regarding the actual affiliations of institutions. Authors are advised to provide their affiliation as indicated on the official internet site of their institution.
Editors
Editorial decisions should not be affected by the origins of the manuscript, including the nationality, ethnicity, political beliefs, race, or religion of the authors. Decisions to edit and publish should not be determined by the policies of governments or other agencies outside of the journal itself.
Human and animal rights
The ethical standards in medical and pharmacological studies are based on the Helsinki declaration (1964, amended in 1975, 1983, 1989, 1996, 2000 and 2013) of the World Medical Association and the Publication Ethics Policies for Medical Journals of the World Association of Medical Journals (WAME).
Authors of studies including experiments on humans or human tissues should declare in their cover letter a compliance with the ethical standards of the respective institutional or regional committee on human experimentation and attach committee’s statement and informed consent; for those researchers who do not have access to formal ethics review committees, the principles outlined in the Declaration of Helsinki should be followed and declared in the cover letter. Patients’ names, initials, or hospital numbers should not be used, not in the text nor in any illustrative material, tables of databases, unless the author presents a written permission from each patient to use his or her personal data. Photos or videos of patients should be taken after a warning and agreement of the patient or of a legal authority acting on his or her behalf.
Animal experiments require full compliance with local, national, ethical, and regulatory principles, and local licensing arrangements and respective statements of compliance (or approvals of institutional ethical committees where such exists) should be included in the article text.
Informed consent
Individual participants in studies have the right to decide what happens to the identifiable personal data gathered, to what they have said during a study or an interview, as well as to any photograph that was taken. Hence it is important that all participants gave their informed consent in writing prior to inclusion in the study. Identifying details (names, dates of birth, identity numbers and other information) of the participants that were studied should not be published in written descriptions, photographs, and genetic profiles unless the information is essential for scientific purposes and the participant (or parent or guardian if the participant is incapable) gave written informed consent for publication. Complete anonymity is difficult to achieve in some cases, and informed consent should be obtained if there is any doubt. If identifying characteristics are altered to protect anonymity, such as in genetic profiles, authors should provide assurance that alterations do not distort scientific meaning.
The following statement should be included in the article text in one of the following ways:
- "Informed consent was obtained from all individual participants included in the study."
- "Informed consent was obtained from all individuals for whom identifying information is included in this article." (In case some patients’ data have been published in the article or supplementary materials to it).
Gender issues
We encourage the use of gender-neutral language, such as 'chairperson' instead of 'chairman' or 'chairwomen', as well as 'they' instead of 'she/he' and 'their' instead of 'him/her' (or consider restructuring the sentence).
Conflict of interest
During the editorial process, the following relationships between editors and authors are considered conflicts of interest: Colleagues currently working in the same research group or department, recent co-authors, and doctoral students for which the editor served as committee chair. During the submission process, the authors are kindly advised to identify possible conflicts of interest with the journal editors. After manuscripts are assigned to the handling editor, individual editors are required to inform the managing editor of any possible conflicts of interest with the authors. Journal submissions are also assigned to referees to minimize conflicts of interest. After manuscripts are assigned for review, referees are asked to inform the editor of any conflicts that may exist.
Appeals and open debate
We encourage academic debate and constructive criticism. Authors are always invited to respond to any editorial correspondence before publication. Authors are not allowed to neglect unfavorable comments about their work and choose not to respond to criticisms.
No Reviewer’s comment or published correspondence may contain a personal attack on any of the Authors. Criticism of the work is encouraged. Editors should edit (or reject) personal or offensive statements. Authors should submit their appeal on editorial decisions to the Editorial Office, addressed to the Editor-in-Chief or to the Managing Editor. Authors are discouraged from directly contacting Editorial Board Members and Editors with appeals.
Editors will mediate all discussions between Authors and Reviewers during the peer review process prior to publication. If agreement cannot be reached, Editors may consider inviting additional reviewers if appropriate.
The Editor-in-Chief will mediate all discussions between Authors and Subject Editors.
The journals encourage publication of open opinions, forum papers, corrigenda, critical comments on a published paper and Author’s response to criticism.
Misconduct
Research misconduct may include: (a) manipulating research materials, equipment or processes; (b) changing or omitting data or results such that the research is not accurately represented in the article; c) plagiarism. Research misconduct does not include honest error or differences of opinion. If misconduct is suspected, journal Editors will act in accordance with the relevant COPE guidelines.
Plagiarism and duplicate publication policy
A special case of misconduct is plagiarism, which is the appropriation of another person's ideas, processes, results or words without giving appropriate credit. Plagiarism is considered theft of intellectual property and manuscripts submitted to this journal which contain substantial unattributed textual copying from other papers will be immediately rejected. Editors are advised to check manuscripts for plagiarism via the iThenticate service by clicking on the "ïThenticate report" button. Journal providing a peer review in languages other than English (for example, Russian) may use other plagiarism checking services (for example, Antiplagiat).
Instances, when authors re-use large parts of their publications without providing a clear reference to the original source, are considered duplication of work. Slightly changed published works submitted in multiple journals is not acceptable practice either. In cases of plagiarism in an already published paper or duplicate publication, an announcement will be made on the journal publication page and a procedure of retraction will be triggered.
Responses to possible misconduct
All allegations of misconduct must be referred to the Editor-In-Chief. Upon the thorough examination, the Editor-In-Chief and deputy editors should conclude if the case concerns a possibility of misconduct. All allegations should be kept confidential and references to the matter in writing should be kept anonymous, whenever possible.
Should a comment on potential misconduct be submitted by the Reviewers or Editors, an explanation will be sought from the Authors. If it is satisfactory and the issue is the result of either a mistake or misunderstanding, the matter can be easily resolved. If not, the manuscript will be rejected or retracted and the Editors may impose a ban on that individual's publication in the journals for a certain period of time. In cases of published plagiarism or dual publication, an announcement will be made in both journals explaining the situation.
When allegations concern authors, the peer review and publication process for their submission will be halted until completion of the aforementioned process. The investigation will be carried out even if the authors withdraw the manuscript, and implementation of the responses below will be considered.
When allegations concern reviewers or editors, they will be replaced in the review process during the ongoing investigation of the matter. Editors or reviewers who are found to have engaged in scientific misconduct should be removed from further association with the journal, and this fact reported to their institution.
Retraction policies
Article retraction
According to the COPE Retraction Guidelines followed by this Journal, an article can be retracted because of the following reasons:
- Unreliable findings based on clear evidence of a misconduct (e.g. fraudulent use of the data) or honest error (e.g. miscalculation or experimental error).
- Redundant publication, e.g., findings that have previously been published elsewhere without proper cross-referencing, permission or justification.
- Plagiarism or other kind of unethical research.
Retraction procedure
- Retraction should happen after a careful consideration by the Journal editors of allegations coming from the editors, authors, or readers.
- The HTML version of the retracted article is removed (except for the article metadata) and on its place a retraction note is issued.
- The PDF of the retracted article is left on the website but clearly watermarked with the note "Retracted" on each page.
- In some rare cases (e.g., for legal reasons or health risk) the retracted article can be replaced with a new corrected version containing apparent link to the retracted original version and a retraction note with a history of the document.
Expression of concern
In other cases, the Journal editors should consider issuing an expression of concern, if evidence is available for:
- Inconclusive evidence of research or publication misconduct by the authors.
- Unreliable findings that are unreliable but the authors’ institution will not investigate the case.
- A belief that an investigation into alleged misconduct related to the publication either has not been, or would not be, fair and impartial or conclusive.
- An investigation is underway but a judgement will not be available for a considerable time.
Errata and Corrigenda
Pensoft journals largely follow the ICMJE guidelines for corrections and errata.
Errata
Admissible and insignificant errors in a published article that do not affect the article content or scientific integrity (e.g. typographic errors, broken links, wrong page numbers in the article headers etc.) can be corrected through publishing of an erratum. This happens through replacing the original PDF with the corrected one together with a correction notice on the Erratum Tab of the HTML version of the paper, detailing the errors and the changes implemented in the original PDF. The original PDF will be marked with a correction note and an indication to the corrected version of the erratum article. The original PDF will also be archived and made accessible via a link in the same Erratum Tab.
Authors are also encouraged to post comments and indicate typographical errors on their articles to the Comments tab of the HTML version of the article.
Corrigenda
Corrigenda should be published in cases when significant errors are discovered in a published article. Usually, such errors affect the scientific integrity of the paper and could vary in scale. Reasons for publishing corrigenda may include changes in authorship, unintentional mistakes in published research findings and protocols, errors in labelling of tables and figures or others. In taxonomic journals, corrigenda are often needed in cases where the errors affect nomenclatural acts. Corrigenda are published as a separate publication and bear their own DOI. Examples of published corrigenda are available here.
The decision for issuing errata or corrigenda is with the editors after discussion with the authors.
COPE Compliance
Terms of Use
This document describes the Terms of Use of the services provided by the Biodiversity Data Journal journal, hereinafter referred to as "the Journal" or "this Journal". All Users agree to these Terms of Use when signing up to this Journal. Signed Journal Users will be hereinafter referred to as "User" or "Users".
The publication services to the Journal are provided by Pensoft Publishers Ltd., through its publishing platform ARPHA, hereinafter referred to as "the Provider".
The Provider reserves the right to update the Terms of Use occasionally. Users will be notified via posting on the site and/or by email. If using the services of the Journal after such notice, the User will be deemed to have accepted the proposed modifications. If the User disagrees with the modifications, he/she should stop using the Journal services. Users are advised to periodically check the Terms of Use for updates or revisions. Violation of any of the terms will result in the termination of the User's account. The Provider is not responsible for any content posted by the User in the Journal.
Account Terms
- For registration in this Journal or any of the services or tools hosted on it, Users must provide their full legal name, a valid email address, postal address, affiliation (if any), and any other information requested.
- Accounts created via this journal automatically sign in the User to the ARPHA Platform.
- Users are responsible for maintaining the security of their account and password. The Journal cannot and will not be liable for any loss or damage from failure to comply with this security obligation.
- Users are solely responsible for the content posted via the Journal services (including, but not limited to data, text, files, information, usernames, images, graphics, photos, profiles, audio and video clips, sounds, applications, links and other content) and all activities that occur under their account.
- Users may not use the service for any illegal or unauthorised purpose. Users must not, in the use of the service, violate any laws within their jurisdiction (including but not limited to copyright or trademark laws).
- Users can change or pseudonomyse their personal data, or deactivate their accounts at any time through the functionality available in the User’s personal profile. Deactivation or pseudonomysation will not affect the appearance of personal data in association with an already published work of which the User is author, co-author, editor, or reviewer.
- Users can report to the Journal uses of their personal data, that they might consider not corresponding to the current Terms of Use.
- The User’s personal data is processed by the Journal on the legal basis corresponding to Article 6, paragraph 1, letters a, b, c and f. of the General Data Protection Regulation (hereinafter referred to as GDPR) and will be used for the purpose of Journal’s services in accordance with the present Terms and Use, as well as in those cases expressly stated by the legislation.
- User’s consent to use the information the Journal has collected about the User corresponds to Article 6(1)(a) of the GDPR.
- The ‘legitimate interest’ of the Journal to engage with the User and enable him/her to participate in Journal’s activities and use Journal’s services correspond to Article 6(1)(f) of the GDPR.
Services and Prices
The Provider reserves the right to modify or discontinue, temporarily or permanently, the services provided by the Journal. Plans and prices are subject to change upon 30 days notice from the Provider. Such notice may be provided at any time by posting the changes to the relevant service website.
Ownership
The Authors retain full ownership to their content published in the Journal. We claim no intellectual property rights over the material provided by any User in this Journal. However, by setting pages to be viewed publicly (Open Access), the User agrees to allow others to view and download the relevant content. In addition, Open Access articles might be used by the Provider, or any other third party, for data mining purposes. Authors are solely responsible for the content submitted to the journal and must confirm [during the submission process] that the content does not contain any materials subject to copyright violation including, but not limited to, text, data, multimedia, images, graphics, photos, audio and video clips. This requirement holds for both the article text and any supplementary material associated with the article.
The Provider reserves the rights in its sole discretion to refuse or remove any content that is available via the Website.
Copyrighted Materials
Unless stated otherwise, the Journal website may contain some copyrighted material (for example, logos and other proprietary information, including, without limitation, text, software, photos, video, graphics, music and sound - "Copyrighted Material"). The User may not copy, modify, alter, publish, transmit, distribute, display, participate in the transfer or sale, create derivative works or, in any way, exploit any of the Copyrighted Material, in whole or in part, without written permission from the copyright owner. Users will be solely liable for any damage resulting from any infringement of copyrights, proprietary rights or any other harm resulting from such a submission.
Exceptions from this rule are e-chapters or e-articles published under Open Access (see below), which are normally published under Creative Commons Attribution 3.0 license (CC-BY), or Creative Commons Attribution 4.0 license (CC-BY), or Creative Commons Public Domain license (CC0).
Open Access Materials
This Journal is a supporter of open science. Open access to content is clearly marked, with text and/or the open access logo, on all materials published under this model. Unless otherwise stated, open access content is published in accordance with the Creative Commons Attribution 4.0 licence (CC-BY). This particular licence allows the copying, displaying and distribution of the content at no charge, provided that the author and source are credited.
Privacy Statement
- Users agree to submit their personal data to this Journal, hosted on the ARPHA Platform provided by Pensoft.
- The Journal collects personal information from Users (e.g., name, postal and email addresses, affiliation) only for the purpose of its services.
- All personal data will be used exclusively for the stated purposes of the website and will not be made available for any other purpose or to third parties.
- In the case of co-authorship of a work published through the Journal services, each of the co-authors states that they agree that their personal data be collected, stored and used by the Journal.
- In the case of co-authorship, each of the co-authors agrees that their personal data publicly available in the form of a co-authorship of a published work, can be distributed to external indexing services and aggregators for the purpose of the widest possible distribution of the work they co-author.
- When one of the co-authors is not registered in the Journal, it is presumed that the corresponding author who is registered has requested and obtained his/her consent that his/her personal data will be collected, stored and used by the Journal.
- The registered co-author undertakes to provide an e-mail address of the unregistered author, to whom the Journal will send a message in order to give the unregistered co-author’s explicit consent for the processing of his/her personal data by the Journal.
- The Journal is not responsible if the provided e-mail of the unregistered co-author is inaccurate or invalid. In such cases, it is assumed that the processing of the personal data of the unregistered co-author is done on a legal basis and with a given consent.
- The Journal undertakes to collect, store and use the provided personal data of third parties (including but not limited to unregistered co-authors) solely for the purposes of the website, as well as in those cases expressly stated by the legislation.
- Users can receive emails from Journal and its hosting platform ARPHA, provided by Pensoft, about activities they have given their consent for. Examples of such activities are:
- Email notifications to authors, reviewers and editors who are engaged with authoring, reviewing or editing a manuscript submitted to the Journal.
- Email alerts sent via email subscription service, which can happen only if the User has willingly subscribed for such a service. Unsubscription from the service can happen through a one-click link provided in each email alert notification.
- Information emails on important changes in the system or in its Terms of Use which are sent via Mailchimp are provided with "Unsubscribe" function.
- Registered users can be invited to provide a peer review on manuscripts submitted to the Journal. In such cases, the users can decline the review invitation through a link available on the journal’s website.
- Each provided peer review can be registered with external services (such as Web of Science Reviewer Recognition Service, formerly Publons). The reviewer will be notified if such registration is going to occur and can decline the registration process.
- In case the Journal starts using personal data for purposes other than those specified in the Terms of Use, the Journal undertakes to immediately inform the person and request his/her consent.
- If the person does not give his/her consent to the processing of his or her personal data pursuant to the preceding paragraph, the Journal shall cease the processing of the personal data for the purposes for which there is no consent, unless there is another legal basis for the processing.
- Users can change/correct their personal data anytime via the functionality available in the User’s profile. Users can request the Journal to correct their personal data if the data is inaccurate or outdated and the Journal is obliged to correct the inaccurate or outdated personal data in a timely manner.
- Users may request the Journal to restrict the use of their personal data insofar as this limitation is not contrary to the law or the Terms of Use.
- Users may request their personal data to be deleted (the right to be forgotten) by the Journal, provided that the deletion does not conflict with the law or the Terms of Use.
- The User has the right to be informed:
- whether his or her personal data have been processed;
- for which purposes the Journal processes the personal data;
- the ways in which his/her personal data are processed;
- the types of personal data that Journal processes.
- The user undertakes not to interfere with and impede the Journal’s activities in the exercise of the provided rights.
- In case of non-fulfillment under the previous paragraph, the Journal reserves the right to delete the user's profile.
Disclaimer of Warranty and Limitation of Liability
Neither Pensoft and its affiliates nor any of their respective employees, agents, third party content providers or licensors warrant that the Journal service will be uninterrupted or error-free; nor do they give any warranty as to the results that may be obtained from use of the journal, or as to the accuracy or reliability of any information, service or merchandise provided through Journal.
Legal, medical, and health-related information located, identified or obtained through the use of the Service, is provided for informational purposes only and is not a substitute for qualified advice from a professional.
In no event will the Provider, or any person or entity involved in creating, producing or distributing Journal or the contents included therein, be liable in contract, in tort (including for its own negligence) or under any other legal theory (including strict liability) for any damages, including, but without limitation to, direct, indirect, incidental, special, punitive, consequential or similar damages, including, but without limitation to, lost profits or revenues, loss of use or similar economic loss, arising from the use of or inability to use the journal platform. The User hereby acknowledges that the provisions of this section will apply to all use of the content on Journal. Applicable law may not allow the limitation or exclusion of liability or incidental or consequential damages, so the above limitation or exclusion may not apply to the User. In no event will Pensoft’s total liability to the User for all damages, losses or causes of action, whether in contract, tort (including own negligence) or under any other legal theory (including strict liability), exceed the amount paid by the User, if any, for accessing Journal.
Third Party Content
The Provider is solely a distributor (and not a publisher) of SOME of the content supplied by third parties and Users of the Journal. Any opinions, advice, statements, services, offers, or other information or content expressed or made available by third parties, including information providers and Users, are those of the respective author(s) or distributor(s) and not of the Provider.
Cookies Policy
Cookies
a) Session cookies
We use cookies on our website. Cookies are small text files or other storage technologies stored on your computer by your browser. These cookies process certain specific information about you, such as your browser, location data, or IP address.
This processing makes our website more user-friendly, efficient, and secure, allowing us, for example, to allow the "Remember me" function.
The legal basis for such processing is Art. 6 Para. 1 lit. b) GDPR, insofar as these cookies are used to collect data to initiate or process contractual relationships.
If the processing does not serve to initiate or process a contract, our legitimate interest lies in improving the functionality of our website. The legal basis is then Art. 6 Para. 1 lit. f) GDPR.
When you close your browser, these session cookies are deleted.
b) Disabling cookies
You can refuse the use of cookies by changing the settings on your browser. Likewise, you can use the browser to delete cookies that have already been stored. However, the steps and measures required vary, depending on the browser you use. If you have any questions, please use the help function or consult the documentation for your browser or contact its maker for support. Browser settings cannot prevent so-called flash cookies from being set. Instead, you will need to change the setting of your Flash player. The steps and measures required for this also depend on the Flash player you are using. If you have any questions, please use the help function or consult the documentation for your Flash player or contact its maker for support.
If you prevent or restrict the installation of cookies, not all of the functions on our site may be fully usable.
Author Guidelines
BDJ uses the highly automated and user-friendly ARPHA Writing Tool (AWT), which navigates you during the authoring and submission process. A Tips & Tricks practical guide to the technicalities within the tool is also available.
The submission process in BDJ starts with creating a manuscript in AWT via the Start new manuscript button. AWT offers a number of fixed yet flexible article template for each article type available in BDJ. Please note that the article templates cannot be changed once the writing process has started.
Select Article Type
Biodiversity Data Journal considers for publication the following categories of papers:
- Data Paper
Formal paper describing large datasets. The article should contain a link to the openly accessible dataset deposited in an internationally recognised repository (see instructions and list of recommended repositories in the Data Publishing Guidelines).
- Omics Data Paper
Description of omics data that can include information about the genome ((meta)genomics), transcription products ((meta)transcriptomics), protein products ((meta)proteomics), and metabolic products ((meta)metabolomics). More information can be found here.
- Methods
Description of novel methodologies and methods used in biodiversity research and biodiversity informatics.
- R Package
Description of an R Package including information on its purpose, installation and usage.
- Software Description
Description of software or an online tool or platform that contains a link to the openly accessible code of the described tool.
- Interactive Key
Online interactive identification key of a given higher taxon or а species group that contains a link to the openly accessible online key.
- Research Article
Publication of a study presenting significant new and original information, other than taxonomic, nomenclatural, faunistic or floristic data. For the latter, please use the "Taxonomy & Inventories" template.
- Taxonomy & Inventories
This manuscript template is designed to encompass various research topics, related to taxonomy, nomenclature, faunistic or floristic explorations and inventories, for example: revision of a group of species or a higher taxon (genus, tribe, family, etc.) comprising nomenclatural and taxonomic novelties, description of new taxa, regional or global species checklists and others. Please note that manuscripts containing such information should use this instead of a Research Article template!
- Species Conservation Profile
A single or multiple IUCN species assessment report(s) imported and edited in an IUCN-compliant species template.
- Alien Species Profile
Assessment report of alien or invasive species following an IUCN-compliant species template. After publication, the article can be integrated with the Global Invasive Species Database (GISD).
- Editorial
Opinion article written by the senior editorial staff.
- Short Communication
Short articles (usually 3-4 pages) addressing preliminary findings of higher interest, unique approaches, controversial opinions, new methods or innovative technologies etc. that demand fast publication.
- Commentary
Short free-text publication meant to draw attention or criticise a previously published work. It does not include original data.
- Forum Paper
Publication meant to initiate or respond to participate in a discussion on a particular scientific topic.
Manuscripts will be accepted for publication only if the following criteria are fulfilled:
- Papers and associated data must contribute to a better understanding of the topic under scrutiny. Studies that have already been published or submissions that are currently under consideration for publication elsewhere will not be accepted for publication.
- Previously published information should be considered and cited in compliance with the good academic practice. References should be complete and accurate. All figures included in manuscripts should be copyright free and duly acknowledge the original source.
- All data underpinning an article, including data tables on which graphs are produced, must be published alongside the paper, e.g. as supplementary files, or links to external repositories where data are deposited, and contain sufficient metadata to facilitate data discovery.
- Manuscripts must be submitted in English - either British/Commonwealth or American English. Authors should confirm the English language quality of their texts or alternatively request thorough linguistic editing prior to peer-review at a price. Manuscripts written in poor English are a subject of rejection prior to peer-review.
- Manuscripts should be concisely written, in a good academic style, and follow a logical sequence. The voice - active or passive - and the tense used should be consistent throughout the manuscript. Results should be clearly and concisely described and supported by the data published with the article, or data published elsewhere but linked to the article.
Materials and Methods
In line with responsible and reproducible research, as well as FAIR data principles, we highly recommend that authors describe in detail and deposit their science methods and laboratory protocols in the open access repository protocols.io.
Once deposited on protocols.io, protocols and methods will be issued a unique digital object identifier (DOI), which could be then used to link a manuscript to the relevant deposited protocol. By doing this, authors could allow for editors and peers to access the protocol when reviewing the submission to significantly expedite the process.
Furthermore, an author could open up his/her protocol to the public at the click of a button as soon as their article is published.
Stepwise instructions:
1. Prepare a detailed protocol via protocols.io.
2. Click Get DOI to assign a persistent identifier to your protocol.
3. Add the DOI link to the Methods section of your manuscript prior to submitting it for peer review.
4. Click Publish to make your protocol openly accessible as soon as your article is published (optional).
5. Update your protocols anytime.
Criteria for Publication
- Originality: Papers and associated data should be sufficiently novel and contribute to a better understanding of the topic under scrutiny. Please consider the examples below to judge if your manuscript might be suitable for BDJ:
- Example 1: Single species occurrence records (e.g., new country or province records) ARE NOT encouraged for submission in BDJ and will most probably be rejected, unless they contain detailed studies and new information on other aspects of species' morphology, genomics, biology, ecology, distribution, etc. (see also Examples 2 and 3 below).
- Example 2: A single species observation must be SIGNIFICANT, either because the species is important in some regard (e.g., medically, or by being an invasive species, or by being endangered, or by being important in biocontrol or biosecurity, etc.), or it expands the species range considerably and represent unexpected discovery of biogeographical or other significance. An additional argument for acceptability of a manuscript could be the presence of images and multimedia (acccompanied by any other new ecological/ethological data), if a species had never been illustrated, or filmed, or recorded for songs. Single species observations are considered novel, if they bring new information based on author(s) personal investigations and do not just repeat or compile already published information.
- Example 3: Multiple species occurrence records are welcome, however occurrence data are considered novel, if they significantly extend the ranges (geographical, temporal or habitat type), or list new country/province records of several species, or concern taxa of high natural or social importance, or feature taxa that are data-poor; occurrence data will NOT be considered novel and suitable for publication, if they only list new localities of a well-known and common (data rich) taxon within a well-studied region.
- Example 4: A local checklist is considered novel, if it includes new data from a locality; a local checklist is NOT considered novel if it is mostly confirmatory and repetitive and lists common species from a locality in a well-studied region.
- Example 5: Stand-alone publications of whole genome or complete mitochondrial genome sequencing are acceptable only if they are part of a complete phylogenetic study and have a significant contribution to the systematics of the taxon.
- Example 6: A data set described in a data paper is considered sufficiently novel if it includes new and mostly unpublished data on a subject, or is a compilation of data on a subject from various appropriately credited, published and/or unpublished sources, or represents new transformations or interpretations of data, including significant data corrections. If the data set is a compilation of data from different sources, all data contributors should be appropriately credited, e.g. through co-authorship or citation, while respecting the use licenses associated with the original data sets. If the data paper describes a dynamic data set (the content of which is expected to change) it is strongly recommended to include links and identifiers in the Data Resources section of the paper to both the dynamic data set and to a data dump deposited in an international archive (e.g. Zenodo). The data dump should reflect the dynamic data as it existed at the time of manuscript submission.
- Data are published: All data underpinning an article, including data tables on which graphs are produced, must be published alongside the paper, e.g. as supplementary files, or links to external repositories where data are deposited, and contain sufficient metadata to facilitate data discovery.
-
Structure: Manuscripts should be concisely written, in a good academic style, and follow a logical sequence. Results should be clearly and concisely described and supported by the data published with the article, or data published elsewhere but linked to the article.
-
Previous research: Previously published information should be considered and cited in compliance with the good academic practice. References should be complete and accurate, where possible including DOIs or links to the article.
Prepare Your Manuscript
Manuscripts for the Biodiversity Data Journal can only be submitted from the online, collaborative, article-authoring ARPHA Writing Tool (AWT) that provides a large set of pre-defined, but flexible, article templates.
To facilitate the writing process, the AWT also provides an automated search and import function from external databases including: electronic registries; catalogues; occurrence data in Darwin Core format; and reference bibliographies.
In the AWT-environment, the authors may invite external contributors, such as mentors, potential reviewers, linguistic and copy editors, colleagues, etc., who are not authors, but may watch and comment on the text during the preparation of the manuscript.
Please consider the following steps, illustrated at the figure below:
- Start a manuscript in the ARPHA Writing Tool (AWT)
- Validate and submit it when ready to the Biodiversity Data Journal
- Make corrections and respond to reviewers and editors completely online
- Submit the revised version at the click of a button
- Work with our copyeditors on final improvements
- Enjoy publication within 3 days after final acceptance
For more information, you may also look at the Editorial Beyond dead trees: integrating the scientific process in the Biodiversity Data Journal and press release The Biodiversity Data Journal: Readable by humans and machines.
Author Contributions
The journal is integrated with Contributor Role Taxonomy (CRediT), in order to recognise individual author input within a publication, thereby ensuring professional and ethical conduct, while avoiding authorship disputes, gift / ghost authorship and similar pressing issues in academic publishing.
During manuscript submission, the submitting author is strongly recommended to select a contributor role for each of co-author, using a list of 14 predefined roles, i.e. Conceptualization, Methodology, Software, Validation, Formal analysis, Investigation, Resources, Data Curation, Writing - Original draft, Writing - Review and Editing, Visualization, Supervision, Project administration, Funding Acquisition (see more). Once published, the article will be including the contributor role for all authors in the article metadata.
Authorship of AI
Preprints
The journal is integrated with the ARPHA Preprints platform, thereby allowing authors to post their pre-review manuscript as a preprint by simply checking the relevant box while completing the submission of their manuscript.
Due to the integration, the authors are not required to re-format or submit any additional files, as the system uses the manuscript to automatically generate a preprint. Subject to a basic editorial screening, the preprint will be posted on ARPHA Preprints within a few days after the manuscript’s submission.
When submitting their manuscripts and requesting a preprint publication authors must keep in mind that preprints are preliminary versions of works accessible electronically in advance of publication of the final version. They are not issued for the purposes of botanical, mycological or zoological nomenclature and are not effectively/validly published in the meaning of the Codes. Therefore, papers containing or dealing with nomenclatural novelties (new names) or other nomenclatural acts (designations of type, choices of priority between names, choices between orthographic variants, or choices of gender of names) will NOT be posted as preprints.
When requesting a preprint publication authors agree that a withdrawal of the published and indexed preprint is only possible if there is a substantial reason for that, i.e. major scientific errors, data fabrication, plagiarism or law infringement. The rejection of the manuscript by the journal is not a valid reason for withdrawal of the preprint. Find out more in this FAQ-section.
Explore the Benefits of posting a preprint or visit ARPHA’s blog to learn more about ARPHA Preprints.
Find more about how to submit your preprint in the ARPHA Manual.
Peer Review
Text and data submitted to this journal will be formally peer reviewed and evaluated for technical soundness and the correct presentation of appropriate and sufficient metadata. All manuscripts undergo a pre-submission technical evaluation in the ARPHA Writing Tool (AWT) environment. The scientific quality and importance of the paper and data will be further judged by the scientific community, through a novel, community-based pre-publication and post-publication peer review.
Reviewers may opt to be anonymous or to disclose their names. The deadlines for the peer review and editorial processes are strict and limited to a maximum of two months after submission.
The peer review process and deadlines described below are articulated on the assumption that the contributions are technically well-prepared and concisely written so that the peer-review is easy, straightforward and not requiring much time from the reviewer.
What is "community peer review"?
It is evident that the peer-review system is increasingly under strain. Our response to this situation is to decrease the load on each individual reviewer without in any way compromising the quality of the final product. The purpose of community peer review is to distribute effort, increase transparency, engage the broader community of experts, and enhance the quality of the science we publish.
Stepwise description of the peer review and editorial process
1. Upon submission, the manuscript is assigned to the Subject Editor responsible for the topic by the in-house Assistant Editor. The Subject Editor is alerted by email.
2. The Subject Editor reads the manuscript and decides if it complies with the journal's scope and should be processed for peer review.
3. The Subject Editor sends review requests to two or three "nominated" reviewers and several other "panel" reviewers.
Note-1: How editors invite reviewers? The journal's database will provide a list of potential reviewers and, if necessary, the editor can add additional names to the list. Review requests will be emailed by a ‘single-click’ button.
Note-2: "Nominated" and "Panel" reviewers. The difference between "Nominated" and "Panel" reviewers is that "Nominated" reviewers are expected to provide a formal review by the deadline; "Panel" reviewers are invited but not required to evaluate the manuscript. Both "Nominated" and "Panel" reviewers can propose changes and corrections, and make comments in the manuscript online and submit a concise reviewer's form.
Note-3: "Community" and "public" peer review. "Community" peer review means that during the review process the manuscript is visible only to the editor, the reviewers and the authors. We are planning to introduce soon an entirely public review process where authors may opt to make their manuscript available for comment by all registered journal users. Reviewers may opt to stay anonymous or disclose their names in either case.
4. The Subject Editor receives a notification email if the nominated reviewer agrees or declines to review the manuscript. In the latter case the editor can appoint alternative reviewers.
5. Reviews are expected within 10 days and can be extended on demand. The Subject Editor will then decide to accept, reject, or request revision of the manuscripts.
Note-4: Provision of reviews. Reviewers will be prompted by automated email notification sent one day before the deadline. In case of delay, the review request can be cancelled automatically, unless an extension has been requested.
6. The authors must provide a revised version of their manuscript within one week, but can ask for an extension, if needed.
7. After submission of the revised version, the Subject Editor compares it against the reviews through an easy-to-use online tool and decides to accept or reject the manuscript. The authors may be asked to make additional revisions, OR in case of substantial changes, the reviewing procedure will be started again.
8. The manuscript will be formatted, proof-read, copy-edited and published within two weeks after acceptance.
Guidelines for reviewers and editors
Reviewers and editors are expected to evaluate the completeness and quality of the manuscript text, related dataset(s), models, workflows or software and their description (metadata), as well as the publication value of data, models, software or workflows. This may include the appropriateness and validity of the methods used, compliance with applicable standards during collection, management and curation of data, and compliance with appropriate metadata standards in the description of the data resources.
The following aspects of evaluation will be considered:
- Originality
- Is the paper sufficiently novel and does it contribute to a better understanding of the topic under scrutiny? Is the work rather confirmatory and repetitive?
- Previous research
- Is previous research adequately incorporated into the paper?
- Are references complete, necessary and accurate?
- Is there any sign that substantial parts of the paper are copied from other works?
- Quality of the manuscript
- Do the title, abstract and keywords accurately reflect the contents and data?
- Is the manuscript consistent, suitably organised and written in grammatically correct English?
- Are the relevant non-textual data and media (data sets, audio and video files, models, software) also available as supplementary files to the manuscript or as links to external repositories?
- Have abbreviations and symbols been properly defined?
- Are conflicts of interest, relevant permissions and other ethical issues addressed in an appropriate manner?
- Quality of the data
- Are the data completely and consistently recorded within the dataset(s)?
- Does the data resource cover scientifically important and sufficiently large region(s), time period(s) and/or group(s) of taxa to be worthy of publication?
- Are the data consistent internally and described using applicable standards (e.g. in terms of file formats, file names, file size, units and metadata)?
- Are the methods used to process and analyses the raw data, thereby creating processed data or analytical results, sufficiently well documented that they could be repeated by third parties?
- Are the data correct, given the protocols? Authors are encouraged to report any tests undertaken to address this point.
- Is the repository to which the data are submitted appropriate for the nature of the data?
- Consistency between manuscript and data
- Does the manuscript provide an accurate description of the data?
- Does the manuscript properly describe how to access the data?
- Are the methods used to generate the data (including calibration, code and suitable controls) described in sufficient detail?
- Is the dataset sufficiently novel to merit publication?
- Does the manuscript put the data resources being described properly into the context of prior research, citing pertinent articles and datasets?
- Have possible sources of error been appropriately addressed in the protocols and/ or the paper?
- Is anything missing in the manuscript or the data resource itself that would prevent replication of the measurements, or reproduction of the figures or other representations of the data?
- Are all claims made in the manuscript substantiated by the underlying data?
Pensoft journals support the open science approach in the peer-review and publication process. We encourage our reviewers to open their identity to the authors and consider supporting the peer-review oaths, which tend to be short declarations that reviewers make at the start of their written comments, typically dictating the terms by which they will conduct their reviews (see Aleksic et al. 2015, doi: 10.12688/f1000research.5686.2 for more details):
Principles of the open peer-review oath
- Principle 1: I will sign my name to my review
- Principle 2: I will review with integrity
- Principle 3: I will treat the review as a discourse with you; in particular, I will provide constructive criticism
- Principle 4: I will be an ambassador for the practice of open science
Post-acceptance Procedure
Frequently Asked Questions (FAQ)
1. Does Biodiversity Data Journal publish only data?
NO! The journal focuses on data, but one can publish analyses and discussions, within the article, as in any other journal.
2. Does Biodiversity Data Journal require all data underlying an article to be published as well?
YES! All small data sets that underpin an article should be imported in the text (e.g., Darwin Core occurrence data, checklists, data tables, literature references) or uploaded as supplemnetary files (e.g., a data table used to create a graph). Large and complex data sets should be deposited in an internationally recognized repository (see Data publication section for details).
3. What kind of data does Biodiversity Data Journal publish?
Any kind of data related to biodiversity, for example: species occurrence data, local or regional checklists, inventories, genomic data, morphological descriptions, ecological observations, environmental data, etc.
4. What is the minimum "publishable" manuscript that can be submitted to the journal?
Any manuscript that brings novel information on any organism from any part of the world. Manuscripts are expected to demonstrate novelty, so it is unlikely that, for instance, a single observation would be sufficient. Please carefully consider our Criteria for publication before you decide to submit a manuscript to BDJ.
5. Why do you define fixed templates for articles?
Templates include some mandatory elements, but they are not fixed. Authors can additional sections or subsections in a manuscript. Using the templates is necessary because the journal's online peer-review and editorial system are designed to automate parts of the publication process to deliver both fast turnover and low cost. These systems do not accept manuscripts written in text processors (e.g., MS Word, or ODT) because they cannot be automated in this way.
6. What? Does Biodiversity Data Journal really NOT accept manuscripts written in MS Word?
NO, it does not! To keep the costs low and affordable for all, manuscripts submitted to the journal must either be written either within a specially designed tool (Pensoft Writing Tool, or PWT) or submitted from integrated external platforms, such as Scratchpads or GBIF Integrated Publishing Toolkit (IPT).
7. What does mean "public", and "community" peer-review?
"Community" peer review means that during the peer-review process the manuscript is visible only to editor, the reviewers and the authors; this is the traditional method in academic publishing and is the default option. Authors may opt, however, to make their manuscript available for comments from all registered journal users ("public" peer review). Reviewers may opt to stay anonymous or disclose their names in either case.
For more information, you may also look at the Editorial Beyond dead trees: integrating the scientific process in the Biodiversity Data Journal and press release The Biodiversity Data Journal: Readable by humans and machines.
8. What is a "Data Paper"?
A "Data Paper" is a scholarly journal publication whose primary purpose is to describe a dataset or a group of datasets, rather than to report a research investigation. As such, it contains facts about data, not hypotheses and arguments in support of the data, as found in a conventional research article. Its purposes are three-fold:
• to provide a citable journal publication that brings scholarly credit to data publishers;
• to describe the data in a structured human-readable form;
• to bring the existence of the data to the attention of the scholarly community.
If you are interested to learn more about it, you may have a look at our Data Publishing page, or the Data Paper Poster.
Data Publishing Guidelines
By submitting to this journal, authors agree to make the data that underpin or are described in their articles publicly available. Authors must include a separate "Data resources" section in their articles, listing datasets and where they are deposited (including accession numbers, DOIs or other persistent URL identifiers). For more information, please see our detailed Strategies and guidelines for scholarly publishing of biodiversity data.
Please remember that publication of data associated with your article in machine-readable form (databases, data sets, data tables) in this journal is mandatory!
The journal will publish papers that strictly adhere to theadhere the rules of the last edition of the International Code of Nomenclature for algae, fungi, and plants. To assure this, authors are requested to follow the instructions below:
Authors of novel fungal taxa must register the new names in only one registry, e.g. either in MycoBank or Index Fungorum but not in both! Please note that the registries regularly coordinate sharing of data and have arranged an informal agreement to only accept the first listed number of any name. Registration of the same new name in multiple registries is considered an inappropriate practice, which creates a considerable confusion and extra work to the registries and necessitates the depreciation of the additional registrations at a later stage!
We provide various modes of data publishing:
- Import of data files in the text (e.g., Darwin Core occurrence data, checklists, specimen and biotic intercations data tables, literature references).
- Supplementary data files (up to 20 MB each) that support graphs, hypotheses, results, etc. published with the article.
- Deposition of large data sets in established thematic or domain-specific international repositories (e.g. GBIF IPT, Dryad, NCBI GenBank, Pangaea, TreeBASE, Morphbank, and others).
- Marked up text published as XML to ensure machine harvesting.
-
A best practice rule and a strong recommendation in this journal is that datasets and software should be deposited and permanently archived in appropriate, trusted, domain-specific repositories that offer long-term preservatiion and persistent identifiers for the deposited datasets (please consult Data Deposition in Open Repositories, or BioSharing, and/or software repositories such as GitHub, GitLab, Bioinformatics.org, or equivalent), which should always be preferred over general ones. The associated persistent identifiers (e.g. DOI, or others) of the dataset(s) must be included in the data or software resources section of the article. Reference(s) to datasets and software should also be included in the reference list of the article with DOIs (where available). Where no domain-specific data repository exists, authors should deposit their datasets in a general repository such as Zenodo, Dryad, Dataverse, or others.
-
Large Darwin Core-structured species occurrence records and observations (primary biodiversity data) should be published with GBIF using either the Integrated Publishing Toolkit (IPT) (for which Pensoft maintains an instance, in case such is not available to the authors - https://ipt.pensoft.net/). Alternatively, DwC data could also be published in trusted and community-recognised repositories (for example, Atlas of Living Australia, Symbiota, Arctos or others), however deposition at GBIF should always have a priority over the alternatives. In case a dataset is deposited in more than one repository, the data paper should link to the dataset which is actually described, again with GBIF having a priority over the others. Authors who want to publish species occurrence data as supplementary files only or through generic repositories (e.g. Zenodo, Dryad), instead of submitting these to GBIF, should justify their decision to do so in a letter to the editors; Small data sets of this kind can be imported into the article text through an Excel template, available in the ARPHA Writing Tool, or via direct online import from GBIF, BOLD, iDigBio and PlutoF (see Import of Darwin Core Specimen Records into Manuscripts).
-
Sampling-event data, that is data describing species occurrences in time and space together with details of sampling effort have to be published through GBIF as well (for details see https://www.gbif.org/sampling-event-data). Such data is available from thousands of environmental, ecological, and natural resource investigations, for example one-off studies or monitoring programmes and many others.
-
Gene sequence and genomic data should be deposited with INSDC (GenBank/EMBL/DDBJ), either directly or via a partnering repository, e.g. Barcode of Life Data Systems (BOLD). Transcriptomics data should be deposited in Gene Expression Omnibus (GEO) or ArrayExpress (see sections Gene Sequence and Genomics for detail).
-
Phylogenetic data should be deposited at TreeBASE.
-
Biodiversity-related geoscience and environmental data should be deposited in PANGAEA.
-
Morphological images other than those presented in the article should be deposited at Morphbank. Images of a specific kind should be deposited in appropriate repositories if these exist (e.g., Morphosource for MicroCT data).
-
Videos should be uploaded to video sharing sites like YouTube, Vimeo or SciVee and linked back to the article text. Similarly, audio files should go to platforms like FreeSound or SoundCloud, and presentations to Slideshare. In addition, multimedia files can also be uploaded as supplementary files on the journal’s website. 3D and other interactive models can be embedded in the article’s HTML and PDF.
-
Other large data sets for which there is no existing thematic or domain-specific repository, could be deposited in the Dryad Data Repository, Zenodo, or Dataverse, either prior to or upon acceptance of the manuscript. This journal is integrated with the Biodiversity Literature Repository (BLR) at Zenodo, where the published article, all images and supplementary data files will be deposited under separate DOIs by the journal upon publication. This does not prevent deposition of other large datasets linked to an article direct at Zenodo by the authors. The journal is integrated also with Dryad, where data should be submitted by the authors. There is a $150 Data Publishing Charge for Dryad submissions, payable via the Dryad website. For more information, please see their FAQ.
-
All external data used in a journal paper must be cited in the reference list, and links to these data (as deposited in external repositories) must be included in a separate data resources section of the article (see How to Cite Data).
For biodiversity and biodiversity-related data the reader may consult the Strategies and guidelines for scholarly publishing of biodiversity data (Penev et al. 2017, Research Ideas and Outcomes 3: e12431. https://doi.org/10.3897/rio.3.e12431). For reader's convenience, we list here the hyperlinked table of contents of these extensive guidelines:
- Data Publishing in a Nutshell
- Data Publishing Policies
- Data Deposition in Open Repositories
- Guidelines for Authors
- Data Published within Supplementary Information Files
- Import of Darwin Core Specimen Records into Manuscripts
- Data Published in Data Papers
- Data Papers Describing Primary Biodiversity Data
- Data Papers Describing Ecological and Environmental Data
- Data Papers Describing Genomic Data
- Software Description Papers
- Guidelines for Reviewers and Editors
Data Quality Checklist and Recommendations
INTRODUCTION
An empowering aspect of digital data is that they can be merged, reformatted and reused for new, imaginative uses that are more than the sum of their parts. However, this is only possible if data are well curated. To help authors avoid some common mistakes we have created this document to highlight those aspects of data that should be checked before publication.
By "mistakes" we do not mean errors of fact, although these should also be avoided! It is possible to have entirely correct digital data that are low-quality because they are badly structured or formatted, and, therefore, hard or impossible to move from one digital application to another. The next reader of your digital data is likely to be a computer program, not a human. It is essential that your data are structured and formatted so that they are easily processed by that program, and by other programs in the pipeline between you and the next human user of your data.
The following list of recommendations will help you maximise the re-usability of your digital data. Each represents a test carried out by Pensoft when auditing a digital dataset at the request of an author. Following the list, we provide explanations and examples of each recommendation.
Authors are encouraged to perform these checks themselves prior to data publication. For text data, a good text editor (https://en.wikipedia.org/wiki/List_of_text_editors) can be used to find and correct most problems. Spreadsheets usually have some functions for text checking functions, e.g. the "TRIM" function that removes unneeded whitespace from a data item. The most powerful text-checking tools are on the command line, and the website "A Data Cleaner's Cookbook" (https://www.datafix.com.au/cookbook/) is recommended for authors who can use a BASH shell.
When auditing datasets for authors, Pensoft does not check taxonomic or bibliographic details for correctness, but we will do basic geochecks upon request, e.g. test to see if the stated locality is actually at or near the stated latitude/longitude. We also recommend checking that fields do not show "domain schizophrenia", i.e. fields misused to containing data of more than one type.
Proofreading data takes at least as much time and skill as proofreading text. Just as with text, mistakes easily creep into data files unless the files are carefully checked. To avoid the embarrassment of publishing data with such mistakes, we strongly recommend that you take the time to run these basic tests on your data.
CHECKLIST
Characters
- The dataset is UTF-8 encoded
- The only characters used that are not numbers, letters or standard punctuation, are tabs and whitespaces
- Each character has only one encoding in the dataset
- No line breaks within data items
- No field-separating character within data items (tab-separated data preferred)
- No "?" or replacement characters in place of valid characters
- No Windows carriage returns
- No leading, trailing, duplicated or unnecessary whitespaces in individual data items
Records
- No broken records, i.e. records with too few or too many fields
- No blank records
- No duplicate records (as defined by context)
Fields
- No empty fields
- No evident truncation of data items
- No unmatched braces within data items
- No data items with values that are evidently invalid or inappropriate for the given field
- Repeated data items are consistently formatted
- Standard data items such as dates and latitude/longitude are consistently formatted
- No evident disagreement between fields
- No unexpectedly missing data
RECOMMENDATIONS
Characters
- The dataset is UTF-8 encoded
Computer programs do not "read" characters like "A" and "4". Instead, they read strings of 0's and 1's and interpret these strings as characters according to an encoding scheme. The most universal encoding scheme is called UTF-8 and is based on the character set called Unicode. Text data should always be shared with UTF-8 encoding, as errors can be generated when non-UTF-8 encodings (such as Windows-1252) are read by a program expecting UTF-8, and vice-versa. (See also below, on replacement characters).
- The only characters used that are not numbers, letters or standard punctuation are tabs and whitespaces
Unusual characters sometimes appear in datasets, especially when databases have been merged. These "control" or "gremlin" characters are sometimes invisible when data are viewed within a particular application (such as a spreadsheet or a database browser) but can usually be revealed when the data are displayed in a text editor. Examples include vertical tab, soft hyphen, non-breaking space and various ASCII control characters (https://en.wikipedia.org/wiki/Control_character).
- Each character has only one encoding in the dataset
We have seen individual datasets in which the degree symbol (°) is represented in three different ways, and in which a single quotation mark (') is also represented as a prime symbol, a right single quotation mark and a grave accent. Always use one form of each character, and preferably the simplest form, e.g. plain quotes rather than curly quotes.
- No line breaks within data items
Spreadsheet and database programs often allow users to have more than one line of text within a data item, separated by linebreaks or carriage returns. When these records are processed, many computer programs understand the embedded linebreak as the end of a record, so that the record is processed as several incomplete records:
item A itemB1 itemC
itemB2
becomes:
itemA itemB1
itemB2 itemC
- No field-separating character within data items (tab-separated data preferred)
Data are most often compiled in table form, with a particular character used to separate one field ("column") from the next. Depending on the computer program used, the field-separating character might be a comma (CSV files), a tab (TSV files), a semicolon, a pipe (|) etc.
Well-structured data keeps the field-separating character out of data items, to avoid confusion in processing. Because commas are commonly present within data items, and because not all programs understand how to process CSVs, we recommend using tabs as field-separating characters (and avoiding tabs within data items!): https://en.wikipedia.org/wiki/Tab-separated_values.
- No "?" or replacement characters in place of valid characters
When text data are moved between different character encodings, certain characters can be lost because the receiving program does not understand what the sending program is referring to. In most cases, the lost character is then represented by a question mark, as in "Duméril" becoming "Dum?ril", or by a replacement character, usually a dark polygon with a white question mark inside.
It is important to check for these replacements before publishing data, especially if you converted your data to UTF-8 encoding from another encoding.
- No Windows carriage returns
On UNIX, Linux and Mac computers, a linebreak is built with just one character, the UNIX linefeed '\n' ('LF'). On Windows computers, a linebreak is created using two characters, one after the other: '\r\n' ('CRLF'), where '\r' is called a 'carriage return' ('CR'). Carriage returns are not necessary in digital data and can cause problems in data processing on non-Windows computers. Check the documentation of the program in which you are compiling data to learn how to remove Windows carriage returns.
- No leading, trailing, duplicated or unnecessary whitespaces in individual data items
Like "control" and "gremlin" characters, whitespaces are invisible and we pay little attention to them when reading a line of text. Computer programs, however, see whitespaces as characters with the same importance as "A" and "4". For this reason, the following four lines are different and should be edited to make them the same:
Aus bus (Smith, 1900)
Aus bus (Smith, 1900)
Aus bus (Smith, 1900)
Aus bus (Smith, 1900 )
Records
- No broken records, i.e. records with too few or too many fields
If a data table contains records with, for example, 25 fields, then every record in the table should have exactly 25 data items, even if those items are empty. Records with too few fields are often the result of a linebreak or field separator within a data item (see above). Records with too many fields also sometimes appear when part of a record has been moved in a spreadsheet past the end of the table.
- No blank records
Blank records contribute nothing to a data table because they contain no information, and a tidy data table has no blank lines. Note, however, that a computer program looking for blank lines may not find what looks to a human like a blank line, because the "blank" line actually contains invisible tabs or whitespaces.
- No duplicate records (as defined by context)
It can be difficult to find duplicate records in some datasets, but our experience is that they are not uncommon. One cause of duplicates is database software assigning a unique ID number to the same line of data more than once. Context will determine whether one record is a duplicate of another, and data compilers are best qualified to look for them.
Fields
- No empty fields
Fields containing no data items do not add anything to the information content of a dataset and should be omitted.
- No evident truncation of data items
The end of a data item is sometimes cut off, for example when a data item with 55 characters is entered into a database field with a 50-character maximum limit. Truncated data items should be repaired when found, e.g.
Smith & Jones in Smith, Jones and Bro
repaired to:
Smith & Jones in Smith, Jones and Brown, 1974
- No unmatched braces within data items
These are surprisingly common in datasets and are either data entry errors or truncations, e.g.
Smith, A. (1900 A new species of Aus. Zool. Anz. 23: 660-667.
5 km W of Traralgon (Vic
- No data items with values that are evidently invalid or inappropriate for the given field
For example, a field labelled "Year" and containing years should not contain the data item "3 males".
- Repeated data items are consistently formatted
The same data item should not vary in format within a single dataset, e.g.
Smith, A. (1900) A new species of Aus. Zool. Anz. 23: 660-667.
Smith, A. 1900. A new species of Aus. Zoologischer Anzeiger 23: 660-667.
Smith, A. (1900) A new species of Aus. Zool. Anz. 23, 660-667, pl. ix.
- Standard data items such as dates and latitude/longitude are consistently formatted
Data compilers have a number of choices when formatting standard data items, but whichever format is chosen, it should be used consistently. A single date field should not, for example, have dates represented as 2005-05-17, May 19, 2005 and 23.v.2005.
- No evident disagreement between fields
If there are fields which contain linked information then these fields should be checked to ensure that they do not conflict with each other. For example, the year or an observation cannot be after the year it was published. Examples:
Year Citation
1968 Smith, A. (1966) Polychaete anatomy. Academic Press, New York; 396 pp.
Genus Subgenus
Aus Bus (Aus)
- No unexpectedly missing data
This is a rare issue in datasets that have been audited, but occasionally occurs. An example is the Darwin Core "verbatimLocality" field for a record containing a full latitude and longitude, but with the "decimalLatitude" and "decimalLongitude" fields blank.
- Spelling of Darwin Core terms
Darwin Core terms are usually considered case sensitive, therefore you should use their correct spelling (http://rs.tdwg.org/dwc/).
We thank Dr. Robert Mesibov for preparing the Data Quality Checklist draft and Dr. Quentin Groom for reviewing it.
FAIR Data Checklist
The present checklist provides guidance for checking on how FAIR your data is. The checklist is adapted from: Jones, Sarah, & Grootveld, Marjan. (2017, November 24). How FAIR are your data? Zenodo. https://doi.org/10.5281/zenodo.1065991 and Preparing fair data reuse and reproducibility. Cornell Research Data Management Service Group.
The answers to all questions with Yes/No are mandatory.
My data and my metadata are Findable.
- My dataset has a persistent identifier, such as a DOI, or other which is cited in the metadata. Yes/No
- The metadata are in an online catalogue or data repository, and can be found there by searching, either with a Web search engine like Google or by using a "Search" function in the catalogue or repository. Yes/No
- Also in the metadata is the recommended format for citing my dataset. Yes/No
My data is Accessible.
- People and computer programs can access my data, although I may put restrictions on that access. Yes/No
- Access is through the persistent identifier and will lead to either my data or the metadata. Yes/No
- A standard protocol (like HTTP) will access my data or metadata, either directly through a Web browser or through an API. Yes/No
My data is Interoperable.
- My data are in a commonly understood and preferably open format, for example, CSV rather than Microsoft Excel's .xlsx. Yes/No
- Where possible, I have used controlled vocabularies or widely accepted formats for all data items, such as ISO 8601 for dates or DarwinCore for specimen records. Yes/No
- The metadata uses machine-readable formats where available, such as ORCID IDs for data authors or, for example, WoRMs APHIA IDs for marine taxa. Yes/No
- Ideally, the metadata is in a machine-readable format such as XML or JSON. Yes/No
My data is Reusable.
- To ensure that my data are understandable by others and can be properly attributed, I have explained clearly in the metadata how, why and by whom the data were created and processed. Yes/No
- The metadata also explains that my data are made openly available with one of the following standard licences:
- Open Data Commons Open Database Licence (https://opendatacommons.org/licenses/odbl/)
- Open Data Commons Attribution Licence (https://opendatacommons.org/licenses/by/)
- Open Data Commons Public Domain Dedication and Licence (https://opendatacommons.org/licenses/pddl/)
- Creative Commons CC-Zero Waiver (https://creativecommons.org/publicdomain/zero/1.0/)
- Creative Commons Attribution Licence CC-BY (https://creativecommons.org/licenses/by/4.0/)
- or with another public copyright licence (https://en.wikipedia.org/wiki/Public_copyright_license).
Linked Data Table for Primary Biodiversity Data
Definition: A linked, or semantically enhanced, data table for primary biodiversity data is a table where data about specimens, taxa, sequences, images, sound recordings, traits, habitat preferences and biotic interactions between specimens or taxa are presented, whenever possible, through hyperlinked persistent unique identifiers, following community-agreed standards and ontologies. The use of hyperlinked identifiers allows for linking to external data resources and also within and between table rows, not only by humans, but also by machines.
This journal strongly encourages authors to present their data in a linked data table, using the template below. The template structure is NOT fixed and can be changed to reflect the specifics and composition of data used in a particular study, following some simple but important rules. The table does not replace or exclude the detailed listing of specimen data (occurrences, label data) in the Materials examined sections of the taxon treatments.
The template does not differ much from what the authors normally use (for example, specimens-BY-sequences-BY-taxa or similar); however, (1) it follows some standardisation in format and structure, (2) it is semantically enriched with links to community-accepted ontologies and external sources of information, (3) information from it can be extracted and distributed to various data aggregators, thus (4) increasing the re-use, discoverability, visibility and citation probability of your research and data (for more details see the blog How to get data from research articles back into the research cycle аt no additional costs).
When filling in the table or changing its structure, please consider the following guidelines:
- The template is read-only. To start working with it, please open and copy it to an editable file in Google Spreadsheet (through the Make a copy option from the File menu) or download it in Excel, CSV, TSV or other format.
- Keep the table as simple and as concise as possible. It is a table of identifiers linked to the actual (meta)data rather than a representation of your complete data.
- The table caption should always start with: "Linked data table of ....".
- Delete all empty columns for which you do not provide data. The table can be reduced to a few columns, depending on the purposes of the study.
- Example-1: If you publish a revision or phylogenetic study of a taxon, it would be sufficient to reduce the table to the first core or source (left) group of data on taxa/specimens, while the entire second target (right) group could be deleted starting with the column of biotic interactions.
- Example-2: If you publish an analysis of the biotic interactions between two groups of taxa, you could reduce the table to: (1) List of source species, (2) Type of interaction, (3) List of target species and (4) Source of information (citation or unpublished source).
- You can add additional columns for data types that are missing in the table. Please do not add columns for data which are already available within the actual data record to which the hyperlinked identifier points to.
- Example: there is no need to add columns for details on localities (e.g. province, district, verbatim descriptions), sampling events, collector names etc., because the link to the Specimen code in GBIF, iDigBio or DiSSCo contains these data in both human- and machine-readable format.
- Changing of the uppermost table headings (rows 1-3) is NOT desirable, because they are linked to ontologies/vocabularies to provide the semantic meaning of data in the respective columns.
- Example-1: if you need to add one more column to list accession numbers of another gene, just copy the previous identical column and paste it to the right, then change the gene name in the heading.
- Example-2: if you need to add a column for a data type which is absent in the template, please name and link it to a term from a community-agreed vocabulary (e.g. Darwin Core, UBERON) or ask us for advice using the email address below.
- Relatively small data tables (e.g. of up to 300-400 records) should be included as an Appendix to the article text and also as a supplementary data file; larger datasets can be deposited at external open thematic repositories and/or supplementary files to the article. In cases where datasets are deposited in an external open repository or as supplementary files to the article, they should be described with a link to the openly available dataset in a Data resources section of the article.
- Most of the hyperlinks to the different data elements in the table will be provided in-house by Pensoft (see the last "Linked by" row in the template). Authors are required to hyperlink only a few data columns for which the linking cannot be provided by the publisher.
- Example: There is no easy way for the publisher to discover and especially to reconcile and link the actual specimen the authors have studied, hence the authors are required to provide these hyperlinks in the table (see "Linked to" row in the template) upon submission of the manuscript.
- Use consistent data formatting when filling in columns, such as Latitude and Longitude, for example (see the journal’s Author guidelines for more details).
- Do not leave fields empty in cases where you have repeatable data for these:
- Example-1: if several specimens of a taxon are listed in the table, please fill in the taxon name in each row (do not list a taxon name only once at its first mention only)
- Example-2: There are still many collections in which specimen data are not indexed in GBIF, iDigBio or DiSSCo (or elsewhere). Please fill in the specimen codes, even when these cannot be hyperlinked to a collection network.
- Controlled vocabulary terms are listed in separate spreadsheets in the document containing the template. The authors are encouraged to select terms from the drop-down menus whenever possible. The examples below explain how the author could add a term missing in a vocabulary or add a new vocabulary in a column, for example, for data types not present in the template, such as Life stage, Body part etc.
- Example-1: In case you would like to fill in a data entity which is not featured in the drop-down menu, you can add an entity as free text in the associated vocabulary spreadsheet (e.g. ‘envo-environments’ for the column ‘Habitat / Environment (after ENVO Ontology)’). This will add your entity to the drop-down menu and you will be able to select it. Please, do not alter any of the other entities in the sheets, as this would prevent us from correctly linking to the ontology terms. For more guidance, you can follow this brief tutorial: https://youtu.be/o96tZZlTs1w.
Video caption: How to add free text to a cell which has a drop-down menu:
- Find the spreadsheet sheet which contains the ontology terms which are listed in the dropdown menu (e.g. ‘envo-environments’ for the column ‘Habitat / Environment (after ENVO Ontology)’) and add your entity as free text at the end of the list in the ‘label’ column. Leave the IRI column empty. You can write a definition of your entity in the ‘definition’ column.
- Go back to the ‘template’ sheet and scroll through the dropdown menu until you find the entity you added. You can now select it.
- Example-2: You can also use the vocabularies which are present in the current document (e.g. uberon-life-stages) or introduce a new vocabulary in a separate spreadsheet and then visualise it as a new data column (please see the tutorial below). https://youtu.be/Byyvhzn7vNw
Video caption: How to add a new column containing a dropdown with ontology terms:
- Insert a new column inside the ‘template’ sheet and name it.
- Go to https://bioportal.bioontology.org/ (or any other ontology repository) to find an ontology that you want to use. BioPortal offers an ontology browser which allows you to search for ontology terms. A similar ontology browser is offered by Ontobee (http://www.ontobee.org/).
- Find an ontology class which describes the types of entities which you want to list in the drop-down menu (e.g. Image class for the different types of images). Copy its resource identifier (ID).
- Go to http://sparql.bioontology.org/, BioPortal’s SPARQL endpoint which allows you to perform semantic queries ontologies featured in BioPortal. Paste in the following query by replacing ‘ID’ with the ID that you copied in the previous step:
PREFIX rdfs: <http://www.w3.org/2000/01/rdf-schema#>
SELECT ?class ?label
WHERE {
?class rdfs:subClassOf <ID>.
?class rdfs:label ?label.
} GROUP BY ?class
Do not forget the angle brackets surrounding the IRI (< >). Click ‘run query’ and copy the results.
- Go back to the spreadsheet containing the template and create a new sheet (in Google Spreadsheets, you can do this from the + button in the bottom menu). Paste the SPARQL query results inside the new sheet. Use the ‘Find and replace’ option to remove any unnecessary characters (e.g. angle brackets (< >) and quotation marks (")). Change the name of the sheet so that it contains the name of the ontology you used and the type of entities you listed.
- Go back to the template sheet and click on the cell where you want the dropdown menu to be. Select ‘Data’ from the upper menu and then ‘Data validation’. For Criteria, select ‘List from range’ and click the table icon to the right to select data range. Go back to the new sheet that you created in step 5 and select the range of values which you want to display in the drop-down menu (e.g. the labels of the different image types). Click ‘Save’ to create the drop-down menu. You can copy the menu along the column rows.
Link to the Google spreadsheet template: https://docs.google.com/spreadsheets/d/1gv7RwKPq7LxGcy114qoEEusU4DLbva3it_7Dh6Nvvt8/edit?usp=sharing
Questions and feedback should be sent to: semanticpublishing@pensoft.net
Data Review Guidelines
Pensoft journals provide data auditing and quality check prior to peer-review, including optional data correcting services, which might be a solution for those authors or their institutions who really care about the quality and re-use of their data. This, however, does not exclude the need for a proper peer review of data, therefore reviewers are kindly asked to look upon some other quality parameters of data published in assoctiation with research articles or as data papers in accordance with the following guidelines:
Quality of the Manuscript
- Does the manuscript conform to the focus and scope of this journal?
- Does the manuscript contain unpublishable — for example fraudulent or pseudoscientific — content?
- Does the manuscript contain sufficiently detailed information to merit publication?
- Do the title, abstract and keywords accurately reflect the contents of the manuscript?
- Is the manuscript internally consistent and suitably organized?
- Is the manuscript written in grammatically and stylistically correct English?
- Are the methods relevant to the study and adequately described?
- Did the authors cite most of the literature pertinent to the subject?
- Are relevant non-textual media (e.g. tables, figures, audio, video) used to an appropriate extent and in a suitable manner?
- Have abbreviations and symbols been properly defined?
- Are the illustrations of sufficient quality?
- Does the manuscript put the data resource being described properly into the context of prior research, citing pertinent articles and datasets?
- Are conflicts of interest, relevant permissions and other ethical issues addressed in an appropriate manner?
Quality of the Data
- Are the data freely and openly available under an appropriate Creative Commons license or waiver?
- Is the repository to which the data are submitted appropriate for the nature of the data?
- Are the data completely and consistently recorded within the dataset(s)?
- Does the data resource cover scientifically important and sufficiently large region(s), time period(s) and/or group(s) of taxa to be worthy of a separate publication?
- Are the data consistent internally and described using applicable standards (e.g. in terms of file formats, file names, file size, units and metadata)?
- Are the methods used to process and analyse the raw data, thereby creating processed data or analytical results, sufficiently well documented that they could be repeated by third parties?
- Are the data plausible, given the protocols? Authors are encouraged to report any tests undertaken to address this point.
Consistency between Manuscript and Data
- Does the manuscript provide an accurate description of the data?
- Does the manuscript properly describe how to access the data?
- Are the methods used to generate the data (including calibration, code and suitable controls) described in sufficient detail?
- Is the dataset sufficiently unique to merit publication as a data paper?
- Are the use cases described in the data paper consistent with the data presented? Would other possible use cases merit comment in the paper?
- Have possible sources of error been appropriately addressed in the protocols and/ or the paper?
- Is anything missing in the manuscript or the data resource itself that would prevent replication of the measurements, or reproduction of the figures or other representations of the data?
- Are all claims made in the manuscript substantiated by the underlying data?
Omics Data Papers
The journal welcomes submission of high-quality data papers describing biodiversity through omic technologies and approaches. The following list illustrates (but is not limited to) the range of data papers that can be considered for publication:
- Units of biodiversity: single genome, single gene studies.
- Biodiversity of populations: population genetics studies.
- Biodiversity at community/ecosystem level: eDNA, environmental monitoring, microbiome.
- Amplicon studies.
In each of these levels of biodiversity, submissions may focus on either or all of the phylogenetic, functional, or metabolic diversity in their systems.
Where do I deposit my omics data and metadata?
In alignment with the FAIR data principles, it is mandatory to deposit datasets used in the manuscript in an external, accessible, and trusted public domain repository.
For sequence based studies, data should be deposited in any of the INSDC databases (ENA, GenBank, DDBJ) and follow the Minimum Information about any (x) Sequence (MIxS) standard.
Data from metabolomic studies should be deposited in any of the member databases of the MetabolomeXchange data aggregation and notification consortium (e.g. EMBL-EBI MetaboLights repository, or the Metabolomics Workbench of NIH).
Proteomics data should be deposited in any of the members of the ProteomeXchange consortium (e.g. the PRoteomics IDEntifications (Pride) Database at the EMBL-EBI, PeptideAtlas, MassIVE, or jPost) and follow the Minimum Information About any Proteomics Experiment (MIAPE) standard. Protein interactions should be deposited to the Database of Interacting Proteins (DIP) or the Biological General Repository for Interaction Datasets (BioGRID), which also accepts datasets of genetic interactions, chemical associations and post translational modifications. Another possible interactions database is the IntAct Molecular Interaction Database.
Functional genomics data can be uploaded to NCBI’s Gene Expression Omnibus (GEO) or EMBL-EBI’s ArrayExpress. Datasets from metagenomic studies should be submitted to MGnify. Genotype and phenotype datasets can be submitted to the European Genome-phenome Archive (EGA), the database of Genotypes and Phenotypes (dbGaP) or the Japanese Genotype-phenotype Archive (JGA).
How do I link my omics data and metadata to the manuscript?
How can I link my code to the manuscript?
Any code specifically generated and used to process the data referenced in each article needs to be reproducible and made available for peer-review. We strongly recommend using R or Jupyter notebooks to combine code with documentation for ease of evaluation and re-use.
Code should be released using a platform such as Github or Code Ocean to facilitate sharing, citing, and evaluation of the code. Dedicated IRIs/URIs/DOIs to the version of the code used in the published version of the manuscript must be included (e.g. the URL to a GitHub release of the manuscript’s code). In addition, authors are encouraged to include compressed archives with the code as supplementary material in case it becomes unavailable through one of the web platforms above in the future.
If analysis or diagnostics were done (in part or in total) using "point and click" software rather than the authors’ own scripts, a very clear and reproducible description of each step must be included in the "Data Processing" subsection of the "Methods" section.
All software used to handle and process the data described in the manuscript must be listed and (where appropriate) cited in the "Technologies used" section of the "Data processing" subsection of the manuscript. All version information, software name and licensing should be included. Authors are not limited by the omics data paper template and can also create a separate section to describe the used software within the "Methods" section or elsewhere.
Automated conversion of omics metadata into data paper manuscript
This journal interoperates with the ENA, a member of the International Nucleotide Sequence Database Collaboration (INSDC) to provide an automated conversion of ENA project metadata into a data paper manuscript in the ARPHA Writing Tool (AWT). After conversion, the manuscript can be further edited and submitted to the journal. Naturally, authors can submit their nucleotide sequence information in any of the INSDC databases, from where data can be linked to the data paper manuscript as described above.
To auto-populate the data paper with metadata and references to the INSDC data, authors should enter the accession number of the ENA study or project (Project ID or Study ID) into the import function of the ARPHA Writing Tool. The tool will import also the associated records from EMBL-EBI’s ArrayExpress and BioSamples databases, if available.
A study or project accession number should be used if all data entries within that project or study (e.g. samples, experiments) are relevant to the data paper. Authors can modify which sequence files are described in the data paper manuscript by adding or deleting resources in the "Data resources" section of the manuscript.
For example, if part of the data described in the data paper manuscript (here in blue) is already uploaded in a project on ENA (e.g. published based on previous publication, here in red), the remaining sequence data will have to be uploaded additionally. Both need to be referenced in the manuscript, specifying the exact sequences used via their resource identifier and URL from where the data can be accessed.
The journal’s import process can automatically harvest and integrate MINSEQE and MIxS compliant metadata which has been submitted to ArrayExpress and BioSamples databases . Information about experimental processes from ArrayExpress is inserted directly in the "Methods section" (not in any particular subsection).
Sample metadata imported from BioSamples is automatically inserted into a supplementary table in a comma-separated value (CSV) format.
If desired, authors are encouraged to edit the original metadata records at the source to ensure that the narrative describes their dataset(s) in sufficient detail. To secure data integrity, deletion and manual re-upload of the supplementary BioSamples table is not permitted. Authors can automatically retrieve the corrected BioSamples records into their manuscript by clicking a button labelled "Re-import from BioSamples" within the Supplementary file editor. If you are using data that is not under your control, please inform the owners of the data for any changes required to ensure data integrity.
Should the metadata not be included in the original submission to one of the repositories listed above, it may be included as a supplementary file to the manuscript, following the community accepted standards for metadata formatting.
Step-by-step guide for importing a data paper manuscript from ENA metadata
The authors of omics data paper manuscripts can automatically import metadata into a manuscript in the ARPHA Writing Tool by following the steps below:
-
Log into the ARPHA Writing Tool and select ‘Create a manuscript’.
-
Select a journal (for now only the Biodiversity Data Journal supports this type of publication) and choose ‘OMICS Data Paper’ from the list named ‘Early research outputs’. Click ‘Import a manuscript’.
-
A new web page containing multiple text fields allows input of various identifiers into different types of papers (Fig. 5). For the omics data paper, insert the respective ENA Study ID or Project ID inside the second text field in the list and click ‘Import’.
-
After a couple of seconds, the imported manuscript will be created. If there are more than 10 samples, associated with the ENA Study, the system would request you to confirm whether all of them should be imported or just the first 10. Regardless of your input, BioSamples metadata about all of the samples would be imported into a supplementary table, if such metadata exists. You can edit any of the article sections or add new ones; you can also add/upload citations, references, figures, supplementary files, etc., and work on the manuscript collaboratively with your co-authors. You can insert additional data resources by clicking a single button, as shown in the figure below. A new data resource object will be generated, along with its respective fields: Download URL, Resource identifier, Data format.
-
If you wish to correct any of the BioSamples records imported into the supplementary table, correct them within the BioSamples database first. Then, click on the "Re-import from BioSamples" button within the Supplementary file editor and the new records would be re-imported.
-
Finally, you should define the usage rights for the described datasets within the "Usage licence" section.
-
After making your modifications to the manuscript, click ‘Validate’ from the menu on the left to ensure that the manuscript is not missing any of the required fields. After that, you can submit the manuscript for technical review. Once the manuscript is confirmed by the editorial office as suitable for the journal, you can further submit it for peer review and publication.
Nanopublications
Nanopublications complement human-created narratives of scientific knowledge with elementary, machine-actionable, simple and straightforward scientific statements that prompt sharing, finding, accessibility, citability and interoperability.
The machine-actionability of nanopublications is a standard due to the fact that each assertion consists of subject, object and predicate (type of relation between the subject and the object), complemented with provenance, authorship and publishing information. The unique feature of the nanopublications is that each of the elements is linked to an online resource, such as controlled vocabularies, ontologies and standards.
They also make it easier for reviewers, scientific editors and, ultimately, readers to focus on and evaluate the very foundations of the study. Additionally, this structured format can also be used to better equip, for example, AI assistants, to correctly ‘read’, ‘interpret’ and deliver the knowledge reported in the publication.
You can find more about the nanopublication workflow and format and its advantages on the Pensoft blog and watch the "Nanopublications for Biodiversity" video.
Within the field-customised workspace available from Nanodash, the authors are guided through an extensive, biodiversity-specialised template, where they can compile the biodiversity association they wish to express, update its provenance and add various information. Conveniently, the templates use domain-specific persistent identifiers (PIDs), in order to link to the widely used and recognised resources, such as ChecklistBank, Catalogue of Life, GBIF, GenBank/ENA, BOLD, Darwin Core, Environmental Ontology (ENVO), Relation Ontology (RO), NOMEN, ZooBank, Index Fungorum, MycoBank, IPNI, TreatmentBank, BioLink.
There are three ways authors in the journal can use nanopublications to future-proof the most important assertions on biological taxa and organisms or statements about associations of taxa or organisms and their environments made in their paper.
Nanopublications associated with a manuscript submitted to BDJ
This workflow lets authors add a Nanopublications section within their manuscript, in order to ‘highlight’ and ‘export’ key points from their papers as nanopublications.
To do this, authors will need to add a Nanopublications section while preparing their manuscript in the ARPHA Writing Tool (AWT). There, in a tabular list, they can underline key statements within a separate pre-designated space. Once a Nanopublication section is added to the manuscript in AWT, the author will need to paste a link to an already existing nanopublication or manage/create it via the Nanodash environment by following a link.
Nanopublications as annotations to existing scientific publications
Nanopublications can also be used by any reader to evaluate and record an opinion about any article using a simple template. The opinion is posted as a nanopublication displayed on the article page (see the Nanopubs tab on the right) and bears a timestamp along with the name of the creator.
To add an annotation, users need to locate the Nanopubs tab in the top-right on any article page and click on the Add reaction command to navigate to the Nanodash workspace accessible to anyone registered on ORCiD. Comments associated with an article - either in-line or article-level comments - can also be exported as nanopublications from the Comment tab.
In both cases, users are navigated to the Nanodash workspace, where they can provide the text of the nanopublication; define its relation to the linked paper using the Citation Typing Ontology (CiTO); update its provenance and add information (e.g. licence, extra creators) by inserting extra elements.
You can find more about the nanopublication workflow and format and its advantages on the Pensoft blog.
How to Access a Manuscript
Manuscripts can be accessed after login
- Login is possible after registration at the journal's website. Our Editorial Office will register all first-time editors and reviewers. New users will receive an automated notification with a request to confirm registration and account information, and options for setting a password, email alerts and other features.
Note: All users can use their registration details to login in all three (Book, E-Book and the respective Journal) platforms of www.pensoft.net.
Note: Please remember that you may have registered with two or more different email addresses, that is why you may have more than one valid account at www.pensoft.net. We advise using only one email address, hence one password associated with it, for all your operations at www.pensoft.net. We highly recommend that in case the user has two or more different accounts, to merge these through the user's profile.
Note: The users can at any time change the initially set password and correct personal details using their user's profile menu (by clicking on the user's name in the upper right corner of the screen appearing after login).
If you have forgotten your password, please use the function Forgot your password? or write to request it from journals@pensoft.net.
There are two ways to access a manuscript
After login, please go to the respective journal’s web page and click on My Tasks button in the upper right corner of the screen. This way, you will be able to see all manuscripts you are responsible for as an author or reviewer or editor.
Note: The manuscripts are grouped by categories, e.g., In Review (no.), In layout (no.), Published (no.), and Archived (no.) etc. The number in brackets after each category shows the number of manuscripts that were assigned to you.
Click on the active manuscript link provided in the email notification you have received from the online editorial system. The link will lead you directly to the manuscript.
General Responsibilities of Editors
Subject, or Associate, editors in Pensoft’s journals carry the main responsibility for the scientific quality of the papers. They take the final decision on a manuscript’s acceptance or rejection and their names are listed as Academic Editor in the header of each published article.
The editorial process is facilitated through an online editorial system and a set of email notifications. The online editorial system informs the Subject Editor about any change in the status of a manuscript from submission to publication.
The online editorial system is designed to save time and effort for Subject Editors in checking the status of the manuscripts. There is no need for editors to visit the journal’s website to keep track on the manuscript they are responsible for. The online system will inform the Subject Editor when an invited reviewer has accepted or declined to review. The email notifications contain stepwise instructions what action is needed at each stage, as well as a link to the respective manuscript (accessible by clicking on the link in the email notification or after login – see How to Access a Manuscript).
Subject Editors are not expected to provide a thorough linguistic editing or copyediting of a manuscript, but rather focus on its scientific quality and overall style, which should correspond to good practices in clear and concise academic writing. It is the author’s responsibility to submit the manuscript in linguistically and grammatically correct English. The Subject Editor should not hesitate to recommend either Reject, or Reject, but resubmission encouraged PRIOR to the peer-review process, in cases when a manuscript is scientifically poor and/or does not conform to journal’s style, and/or is written in poor English (see Note under point 1 below how to reject a manuscript prior to peer review).
Editors-in-Chief, Managing Editors or their deputies are allowed to publish a limited proportion of papers per year co-authored by them, after considering some extra precautions to avoid an impression of impropriety, endogeny, conflicts of interest and ensure that the editorial decision-making process is transparent and fair.
It often happens that even carefully written manuscripts may contain small errors in orthography or stylistics. We shall be thankful if editors spot such errors during the reading process and correct them.
Stepwise Description of the Editorial Process
Once a manuscript is submitted, the Managing Editor (or the Editor-in-Chief) briefly checks the manuscript for any personal notes addressed to the E-i-C during the submission process, any explicitly indicated conflict of interest or request for additional services or discount and then the Editor can process it for review.
Note: The workflow that the journal uses allows the Managing Editor (or the Editor-in-Chief) to perform a thorough initial check of the manuscript, during a previous stage called Technical evaluation, prior to its submission to the journal. During this stage the Editor checks the manuscript for conformance with the journal's Focus, Scope, Policies and style requirements and decides whether it is potentially suitable for publication and can be processed for review. In case the submission does not comply with the journal's standards, it can be either 1) sent back to the authors for correction - this process can be iterated in as many rounds as needed - or 2) rejected in ARPHA, i.e. disabled for submission to the journal. Either way, the system requires that the Editorial office add a note explaining the reason for return or rejection.Once a manuscript has been approved during the Technical evaluation and then submitted to the journal, the Managing Editor (or the Editor-in-Chief) assigns it to the Subject Editor responsible for the respective topic (e.g., science branch or taxon). The Subject Editor receives a notification email on the assignment.
Note: The link to the respective manuscript is available in the review request email and all consequent reminder emails. The manuscript is accessible by clicking on the link in the email notifications, or via the user's dashboard after login. Please see How to Access a Manuscript above in case you have any difficulties.The Subject Editor has to read the manuscript and decide whether it is potentially suitable for publication and can be processed for review, or rejected immediately, or returned to the author for improvement and re-submission. Reasons for rejection can be a low scientific quality, non-conformance to the journal’s style/policies, and/or linguistically or grammatically poor English language.
Note: There are two ways to reject a manuscript prior to review process:
- Through the buttons Reject or Reject, but re-submission encouraged in the Editorial tab. Please note, however, that the buttons will be made active only after a justification for the rejection is provided in the text field.
- Through an email to the Editorial office explaining the reason for rejection. The manuscript will be then rejected/returned through the online editorial system and the respective notification email will be sent from the Editorial Office.In case the manuscript is acceptable for peer review, the Subject Editor has to invite reviewers by clicking on the Invite reviewers link. A list of reviewers will appear from which the editor can choose the appropriate ones or add new.
Note: "Nominated" and "Panel" reviewers. The difference between "Nominated" and "Panel" reviewers is that "Nominated" reviewers are expected to provide a formal review by the deadline; "Panel" reviewers are invited but not required to evaluate the manuscript. Both "Nominated" and "Panel" reviewers can propose changes and corrections, and make comments in the manuscript online and submit a concise reviewer's form.Once reviewers are chosen, the Subject Editor has to click the Invite reviewers green button at the end of the page which will generate emails templates with review invitations. It is highly recommended that the Subject Editor adds some personal words above the standard email text review invitation.
In case a reviewer is absent from our users' data base, the editor can add his/her name and email through the Add new reviewer link, which will appear once the search field reveal no results. It is possible that the needed reviewer has already been registered in the Pensoft database either as customer or author/reviewer of another journal. If this is the case, then his/her name, affiliation and other metadata will automatically appear once the e-mail field is populated in the Create user online form.
The Subject Editor receives a notification email if the reviewer has agreed to review a manuscript or declined to do that. The editor takes care to appoint additional reviewers in case some of the invited reviewers have declined.
Once all reviewers submit their reviews, the Subject Editor receives an email notification, inviting him/her to consider reviewers' opinions, read through the manuscript and make a decision through the Proceed button. The Subject Editor is also notified in case none of the reviewers agreed to provide a formal review on the manuscript and he/she is prompted to consider inviting more reviewers.
Note: Editorial comments should be added in the online editorial form; comments and corrections applied using the Track Changes feature and these will be consolidated in a single document along with all reviewers' comments and suggestions.At this stage, the editor should make a decision either to (1) accept the manuscript, or (2) reject it, or (3) initiate another review round. In case the manuscript is not rejected, but recommended for Minor Revision, Major Revision, or Acceptance, the author is expected to submit a revised version within a certain period of time and the Subject Editor will be notified by email about that.
Note 1: Authors must submit revised versions using the Track Changes feature and resolve all Comments, replying to each Subject Editor and reviewer's query so that the Subject Editor can see their corrections/additions. Authors are expected to reply to the essential critiques and comments of reviewers separately through the online editorial system.
Note 2: During the second, or next, review round, the Subject Editor may decide to ask reviewers to evaluate the revised version of the manuscript. He/she may also make a decision based on the author’s responses and the revised version of the manuscript without asking additional reviewers' support.After acceptance, the manuscript will go to layout and proofreading. The Subject Editor will be notified by email when the final proof is uploaded on the journal’s website. The Subject Editor is expected to look at the proofs and notify the Editorial Office through email in case the proofs need improvement.
The Subject Editor may always access information on the manuscripts which have been edited by him/her through the menu My Tasks –> Subject Editor on the journal’s web page – In Review (no.), In Edit (no.), Published (no.), and Archived (no.). The number in brackets after each category shows the number of manuscripts that were assigned.
Editors’ and Reviewers’ Workload Stats
While selecting a Reviewer or a Subject Editor to assign to a manuscript, Editors can access the current and past workload for the person they are considering.
By clicking on the user’s name, an Editor sees how many editorial or review tasks the person is currently assigned with, as well as a record of the user’s previous performance across all ARPHA-hosted journals (i.e. number of accepted and declined editorial and review assignments, as well as the titles of the corresponding journals).
The feature is meant to facilitate and expedite the editorial process by discouraging assignment of tasks to overburdened or inactive users.
Find how to Manage Subject editor assignments and Invite Reviewers in the ARPHA Manual.
Review Quality Rating
Subject Editors should evaluate each review submitted to a manuscript they are handling by using a 5-star rating system. The average score is visible for Subject editors who consider the user as a Reviewer. The feature is meant to expedite the editorial process by aiding Subject Editors in the selection of the most suitable reviewers.
Find how to Rate a peer review in the ARPHA Manual.
Guidelines for Reviewers
Pensoft journals support the open science approach in the peer review and publication process. We encourage our reviewers to open their identity to the authors and consider supporting the peer review oaths, which tend to be short declarations that reviewers make at the start of their written comments, typically dictating the terms by which they will conduct their reviews (see Aleksic et al. 2015, doi: 10.12688/f1000research.5686.2 for more details):
Principles of the open peer-review oath
- Principle 1: I will sign my name to my review
- Principle 2: I will review with integrity
- Principle 3: I will treat the review as a discourse with you; in particular, I will provide constructive criticism
- Principle 4: I will be an ambassador for the practice of open science
How to Access a Manuscript
Manuscripts can be accessed after login
- Login is possible after registration at the journal's website. Our Editorial Office will register all first-time editors and reviewers. New users will receive an automated notification with a request to confirm registration and account information, and options for setting a password, email alerts and other features.
Note: All users can use their registration details to login in all three (Book, E-Book and the respective Journal) platforms of www.pensoft.net.
Note: Please remember that you may have registered with two or more different email addresses, that is why you may have more than one valid account at www.pensoft.net. We advise using only one email address, hence one password associated with it, for all your operations at www.pensoft.net. We highly recommend that, in case the user has two or more different accounts, to merge these through user's profile.
Note: Users can at any time change the initially set password and correct personal details using their user's profile menu (by clicking on the user's name in the upper right corner of the screen appearing after login). If you have forgotten your password, please use the function Forgot your password? or write to request it from journals@pensoft.net.
There are two ways to access a manuscript
After login, please go to the respective journal’s web page and click on My Tasks button in the upper right corner of the screen. This way, you will be able to see all manuscripts you are responsible for as Author or Reviewer or Subject Editor.
Note: The manuscripts are grouped by categories, e.g., In Review (no.), In layout (no.), Published (no.), and Archived (no.) etc. The number in brackets after each category shows the number of manuscripts assigned to you.
Click on the active manuscript link provided in the email notification you have received from the online editorial system. The link will lead you directly to the manuscript.
General Responsibilities of Reviewers
This journal uses a single-blind peer review process. The reviewers are encouraged to disclose their identity, if they wish so. The peer review and editorial process is facilitated through an online editorial system and a set of email notifications. The online editorial system sends the Reviewer a review request, initiated by the Subject Editor or the Editorial Office. The online system will also inform about delays in the reviewing and will confirm a successful review submission. The email notifications contain stepwise instructions about the actions needed at each stage along with the link to the respective manuscript (accessible only after login – see section How to Access a Manuscript).
Reviewers are not expected to provide a thorough linguistic editing or copyediting of a manuscript, but rather focus on its scientific quality and overall style, which should correspond to the good practices in clear and concise academic writing. If Reviewers recognize that a manuscript requires linguistic edits, we shall be grateful for them to inform both the Author and the Subject Editor in the report. It is the Author’s responsibility to submit the manuscript in linguistically and grammatically correct English.
It often happens that even carefully written manuscripts may contain small errors in orthography or stylistics. We shall be thankful if Reviewers spot such errors during the reading process and correct them.
The manuscripts will generally be reviewed by two or three experts with the aim of reaching a first decision as soon as possible. Reviewers do not need to sign their reports, but are welcome to do so. They are also asked to declare any conflicts of interest.
Reviewers are asked whether the manuscript is scientifically sound and coherent, how interesting it is and whether the quality of the writing is acceptable. Where possible, the final decision is made on the basis of the peer reviews. In cases of strong disagreement between the reports or between the authors and peer reviewers, the editor can assess these according to his/her expertise or seek advice from a member of the journal's Editorial Board.
The ultimate responsibility for editorial decisions lies with the respective Subject Editor and/or, in some journals, with the Editor-in-Chief. All appeals should be directed to the Editor-in-Chief, who may decide to seek advice from the Subject Editors or the Editorial Board.
Reviewers are also asked to indicate which articles they consider to be especially interesting or significant. These articles may be given greater prominence and greater external publicity, including press releases addressed to science journalists and mass media.
During a second review round, reviewers may be asked to evaluate the revised version against their recommendations submitted during the first review round.
Reviewers are kindly asked to be polite and constructive in their reports. Reports that may be insulting or uninformative will be rescinded.
Reviewers are asked to start their report with a very brief summary of the reviewed paper. This will help the editor and the authors see whether the reviewer correctly understood the paper or whether a report might be based on misunderstanding.
Furthermore, reviewers are also asked to comment on originality, structure and previous research:
Originality: Is the paper sufficiently novel and does it contribute to a better understanding of the topic under scrutiny? Is the work rather confirmatory and repetitive?
Structure: Is the introduction clear and concise? Does it place the work into the context that is necessary for a reader to comprehend aims, hypotheses tested, experimental design or methods? Are Material and Methods clearly described and sufficiently explained? Are reasons given when choosing one method over another one from a set of comparable methods? Are the results clearly, but concisely described? Do they relate to the topic outlined in the introduction? Do they follow a logical sequence? Does the discussion place the paper in scientific context and go a step beyond the current scientific knowledge on the basis of the results? Are competing hypotheses or theories reasonably related to each other and properly discussed? Do the conclusions seem reasonable?
Previous research: Is previous research adequately incorporated into the paper? Are references complete, necessary and accurate? Is there any sign that substantial parts of the paper are copies of other works?
Stepwise Description of the Peer Review Process
-
This journal uses a single-blind peer review process. Notwithstanding with that, the reviewers are encouraged to disclose their identities, if they wish to do so.
-
The Reviewer receives a review request generated by the Subject Editor or the Editorial Office and is expected to either agree to provide a review, or decline, through pressing the Will do the review or Unable to do the review link in the online editorial system. In case the Reviewer agrees to review the manuscript, he/she should submit the review within a certain time frame, which may vary in the different journals.
Note: The link to the respective manuscript is available in the review request email and all consequent reminder emails. The manuscript is accessible by clicking on the link in the email notification, or after login. Please look at the section How to Access a Manuscript above in case you have any difficulties. -
The review should be submitted through the Proceed button. The review should consist of (1) a simple online questionnaire to be answered by ticking either Yes, No, or N/A; and (2) comments addressed to both the Author and the Editor in the online form.
Note: Reviewers can insert comments in the manuscript review version. -
The Reviewer may decide to stay anonymous or open his/her identity by ticking the Show my name to the author(s) box at the bottom of the reviewer’s form.
-
The review process is completed by selecting a recommendation from five options: (1) Reject; (2) Reject, but resubmission encouraged; (3) Major Revision; (4) Minor Revision; (5) Accept. The system will ask for one more confirmation of the selected recommendation before submission. The submitted review cannot be changed after submission.
Note: Reasons for rejection can be a low scientific quality, non-conformance to the journal’s style/policies, and/or grammatically poor English language. -
Once a Reviewer submits a review of a manuscript, he/she receives an acknowledgement email from the journal.
-
The submission of the review is also automatically reported to Clarivate - Web of Science Reviewer Recognition Service (formerly Publons). Reviewers are asked to confirm whether they want their reviews to be recorded on Clarivate.
-
When all Reviewers have submitted their reviews, the Subject Editor makes a decision to either accept, reject or request further minor/major revision.
-
After Subject Editor's decision, the manuscript is sent back to the author for comments and further revision. The author needs to submit a revised version in due time.
-
Reviewers are notified via email when the revised version of a manuscript they have reviewed is submitted by the author. They receive a link to the revised version along with the editorial decision and all reviews of the manuscript. Reviewers are also provided with a feedback form should they have any comments on the revised version.
-
When an article is published, all Reviewers who have provided a review for the respective manuscript receive an email acknowledgement. In the email, there is a link to view/download the published article.
-
The Reviewer may always access information on the manuscripts that are being / have been reviewed by him/her through the menu My Tasks –> Reviewer on the journal’s web page – In Review (no.), In Edit (no.), Published (no.), and Archived (no.). The number in brackets after each category shows the number of manuscripts that have been assigned to you.
Benefits for Editors and Reviewers
This journal does not exclude editors from publishing papers in the journal (co-)authored by them. However, this is only possible for a limited proportion of papers per year, with some extra precautions and procedures to avoid an impression of impropriety, endogeny or conflicts of interest, and to ensure the editorial decision-making process is transparent and fair. For more information please consult the Policies page on the journal's website.
Pensoft editors and reviewers are entitled to a set of benefits in appreciation for their contribution to the quality of the works we publish. Please make sure to apply for your discount prior to the manuscript submission.
|
For Editors |
For Reviewers |
|
|
* When an individual qualifies for multiple discounts Pensoft will use the largest that applies.
Topical collections
Topical collections are collections of articles grouped together by a shared topic or interest group, such as an emerging area of research, proceedings of a conference, outcomes of a research project or a Festschrift volume.
Article collections aim to aid the dissemination and outreach of multiple research outcomes and also bring together research teams from around the globe working on similar topics, thus increasing the opportunities for collaboration, sharing and re-use of research. Article collections bring credit, increased discoverability, visibility and recognition to both their collection editors and participating authors.
Topical collections can be opened in any journal hosted on ARPHA and can be permanent or made subject to a submission deadline. It is only up to the Collection editor(s) to decide whether and when the collection is to be closed for submission (given a timely public announcement is provided). The articles are published on a rolling basis, as soon as they are ready for publication, and can be part of different journal issues, published across many years.
Article collections are managed by a Collection editor and associated Guest editors. To pitch a Topical collection, either contact the Editor-in-Chief or submit an Open an article collection proposal form. Before pitching a Topical collection, please make yourself aware of the specificity of the focus, scope and policies of the journal and the associated responsibilities and benefits for you as a Collection editor.
How It Works
The following guidelines apply for Topical collections in the ARPHA journals.
-
Article collections can be opened in any of the ARPHA-hosted journals. It is subject to the journal's policy, however, to offer this feature or not.
-
Collections may have subcollections, for example, topical subcollections. A subcollection cannot be managed separately from the parent collection, except in the case of conference proceedings submission workflows available at some ARPHA-hosted journals.
-
Opening and managing a collection
-
The article collections are managed by a Collection editor and Guest editors. The Collection editor is responsible for approving or declining manuscripts submitted to the article collection; assigning a Guest editor to each manuscript for handling the peer review process; and managing the collection on the journal’s website (e.g. change the collection’s description or the order of the papers). The Collection editor has the full rights of a Guest editor and can also handle manuscripts.
-
Before pitching a collection, assure that you are ready to appoint other Guest editors, if necessary. The Collection editor and the Guest editors are also expected to commission an initial set of manuscripts to be submitted soon after the opening of the collection.
-
Submit the Article collection proposal form or contact the Editor-in-Chief via email. The collections should fully comply with the journal’s focus, scope and editorial policies.
-
Online proposals are forwarded to the Editor-in-Chief and to the journal’s editorial office for approval. The editorial office checks and confirms the guest editors' credentials.
-
Upon approval of the proposal, the journal’s editorial office will set up the collection on the journal’s website.
-
Open collections will be promoted through the journal’s website and social media in collaboration with the Collection and Guest editors.
-
Editors of Topical collections with no set submission deadlines need to inform the journal’s editorial office if they wish to close the collection for submissions in a timely manner.
-
-
Authors opt for assigning their manuscript to a collection during submission. In case the manuscript is declined from the collection, it undergoes the regular evaluation and peer review process at the journal.
-
Once the manuscript passes the initial pre-review screening by the Editor-in-Chief and the journal's editorial office, it is forwarded to the Collection editor to either approve or decline it for the collection. The Collection editor is notified about each new submission to the collection via email sent by the system.
-
After reading the paper, the Collection editor can:
-
accept it in the collection and assign it to a Guest editor.
-
decline it from the collection and send it back to the journal's editorial office.
-
-
Once a manuscript is assigned to a Guest editor, he or she takes on the responsibility to invite reviewers and provide an editorial decision for revision, rejection or acceptance of the manuscript, based on the reviews and personal evaluation. Papers submitted by the guest editor(s) must be handled under an independent review process and make up no more than 25% of the collection's total.
-
The editorial decisions are automatically forwarded to the authors by the system.
-
The guest editors are overseen by the journal's Editor-in-Chief and/or dedicated board members, and may intervene in the editorial process. Depending on the journal’s policy, the journal’s Editor-in-Chief might need to approve the Guest editor’s final decision before the manuscript is accepted for publication.
Benefits of Editing a Collection
The main advantages to open and edit an article collection can be summarised as follows:
Credit and recognition for the Collection and Guest editors who take care to organise and manage the article collection.
Facilitates discoverability and usability of topically related studies, which in turn benefits both authors and readers.
Increases the visibility of related papers, even when papers might otherwise lack in viewership.
Prompts simultaneous citation of multiple articles related to a certain subject.
Facilitates citation and referencing of the whole issue as a complete entity.
To show our gratitude to the collection editors, we are also providing a free publication to collection editors in the collection they edit and manage.
Editor’s Responsibilities
By proposing an article collection (Topical collection), you agree to act as a Collection editor, whose main responsibilities are:
Working with the editorial office to set up the article collection on the journal’s website.
Appoint Guest editors for the article collection.
Approve or decline each manuscript submitted to the article collection.
Assign a Guest editor for each manuscript submitted to the article collection.
Assure that the article collections complies with any relevant requirements, as set up by the journal and the agreement (if any).
Inform the journal’s editorial office about any changes or issues concerning the management of the collection in due time.
You will also be granted the user rights of a Guest editor necessary to handle manuscripts in the system (i.e. assign reviewers and provide an editorial decision on the acceptance/rejection of the manuscript).
The responsibilities of a Guest editor are:
Handling the peer review of the manuscripts they have been assigned to.
Making an editorial decision for revision, acceptance or rejection of the manuscripts they have been assigned to, based on the reviews provided and personal evaluation.
Taking into consideration the recommendations of the journal’s Editor-in-Chief.
For more information about the editorial workflow, visit How it works?
Publisher's statement
A key policy and strategic aim of Pensoft is to provide high-quality and inclusive publishing services at highly competitive and affordable Article Processing Charges (APCs) or for free through its diamond open access journals. See Pensoft’s journal portfolio here.
In order to ensure long-term sustainability of the journals and cover the cost of the associated in-house publishing services, our journals require Article Processing Charges (APCs). These charges apply only after a submitted manuscript is accepted for publication, and may be partially or fully covered by institutional funds to reduce financial burdens on authors of research.
Pensoft strongly supports measures that ensure an inclusive and FAIR publishing environment, which in turn prompts quality, sustainability and reasonable pricing in scholarly publishing. You can find more about the publisher’s view on quality, transparency, openness and equity in scholarly publishing in Pensoft’s official statement, prompted by the publication of the European Union’s Conclusions on high-quality, transparent, open and equitable scholarly publishing.
In compliance with the Plan S requirements, Pensoft provides a breakdown of the APC following the guidelines by the Fair Open Access Alliance (FOAA). The report on the journal’s APC is submitted on a yearly basis to the Journal Comparison Service by Coalition S and the detailed breakdown is available to the participating funding institutions on the platform.
Authors who are unable to pay their APCs for several reasons, should consult the Journal’s Discounts and Waivers page, use the diamond open-access journals (free to publish and free to read) hosted on Pensoft’s ARPHA Publishing Platform, or contact the journal’s Editor-in-Chief directly.
Core Charges
Core services included in our Article Processing Charges:
- Manuscript authoring in the ARPHA Writing Tool
- Online collaboration with your co-authors and peers during authoring
- Online import of occurrence records into manuscripts from GBIF, BOLD, iDigBio and PlutoF
- Automated creation of data paper manuscript from GBIF IPT, DataONe EML and BEXIS files
- Online search and import of cited references and data from CrossRef, DataCite, PubMed, RefBank, GNUB, and Mendeley
- Automated technical check for consistency of the manuscript
- Pre-submission technical and editorial checks by the editorial office
- Pre-submission peer-review, organized by the author (optional)
- Pre-publication peer-review
- Community-driven post-publication peer-review (optional)
- Automated registration of peer reviews at Clarivate (formerly Publons)
- Registration of new taxa in ZooBank and IPNI
- Export of occurrence records in Darwin Core Archive to GBIF
- Export of taxon treatments to EOL, Plazi and Species-ID
- Export of taxon treatments in Darwin Core Archive
- Publication in semantically enhanced HTML, PDF and JATS XML formats
- Machine-readable, harvestable content via JATS XML and Web services
- Archiving in trusted international repositories
- Active dissemination via social networks and email alerts
Please note that the charges below are applicable for all manuscripts submitted after 1st of January 2022. Papers of special interest to the journal and reviews of special importance for science are to be priced by agreement.
|
Article type |
Size limits* |
Article processing charges |
|
Character count including spaces |
||
|
Editorial |
10,000 |
By invitation only |
|
Corrigendum** |
5,000 |
€ 100 |
|
All other*** |
40,000 |
€ 650 + € 10 for each 1000 characters above 40,000 |
* Please ask for a quote for manuscripts that exceed the indicated limit more than two times.
**Corrigenda to papers published in this journal.
***Given the importance of publishing occurrence records (materials examined) in both human- and machine-readable formats within the article text, discounts are negotiable for large papers containing a lot of this type of data.
Please note that the above prices do not include VAT (Value Added Tax). VAT is applicable only for VAT NON-registered customers based within the European Union. To avoid charging VAT, the EU companies or persons should provide their VAT registration numbers validated with the EU taxation database (https://ec.europa.eu/taxation_customs/vies/).
Article Collections
Article collections enable conference organizers or project coordinators to publish a number of articles under a common theme and editorship. Depending on the number of articles to be included, Pensoft offers discounts on APCs as described in the table below.
|
|
Small |
Medium |
Large |
|
Number of articles |
< 10 |
10 – 20 |
21 + |
|
Discount on APCs |
5% |
10% |
15% |
|
PR campaign |
By agreement |
By agreement |
Included |
|
Institutional branding |
By agreement |
By agreement |
Included |
We are happy to discuss alternative arrangements if there is a better way to suit your needs for n article collection. Please do not hesitate to contact us or to submit your proposal through the article collection application form.
Discounts and Waivers
Please note that the discounts and waivers policy below is applicable for all manuscripts submitted after 1st of January 2024.
Authors can apply for a discount or a waiver during manuscript submission if they comply with the conditions listed below. The journal will not consider requests made during the review process or after acceptance. Formal letters to the editors will not be considered outside the application process during manuscript submission. The waiver system will be managed by administrative staff not involved in decisions regarding article acceptance. We ask authors not to discuss any issues concerning payment with editors.
- Discount of 10 % is offered to:
- Scientists working privately, not affiliated with an institution.
- Graduate and PhD students if they are first authors of a manuscript.
- Discount of 15 % is offered to:
- Scientists affiliated with institutions located in Research4Life Group B countries (https://www.research4life.org/access/eligibility/#groupb) if they are lead or corresponding authors of a manuscript. In cases of multiple affiliations, all institutions should be located in eligible countries.
- Discounts are also offered to our editors and reviewers, for more information see here.
- Special discounts can be requested by the authors of extensive review papers and monographs.
- Waivers (once per year per (co-) author for manuscripts no larger than 10 printed pages, or for the first 10 pages of a larger manuscript) are offered to:
- Retired scientists who are editors or active reviewers for this journal (1-3 reviews provided in the year before the manuscript submission).
- Scientists affiliated with institutions located in Research4Life Group A countries (https://www.research4life.org/access/eligibility/#groupa), if they are lead or corresponding authors of a manuscript. In cases of multiple affiliations, all institutions should be located in eligible countries.
The journal offers also various institutional programs and membership plans to support Open Access scientific publishing. To be eligible, the author must be a corresponding author affiliated with the institution or agency.
Discounts and waivers do not accumulate.
Please note that the discounts and waivers policy below is applicable for all manuscripts submitted before 1st of January 2024.
Authors can apply for discount or waiver during manuscript submission if they comply with the conditions listed below. The journal will not consider requests made during the review process or after acceptance. Formal letters to the editors will not be considered outside the application process during manuscript submission.
- Discount of 10 % is offered to:
- Scientists working privately, not affiliated with an institution.
- Graduate and PhD students if they are first authors of a manuscript.
- Scientists living and working in lower middle-income countries (http://data.worldbank.org/income-level/lower-middle-income) if they are sole authors of a manuscript, or authors' research is funded primarily (50% or more of the work contained within the article) by an institution or organization from the eligible countries.
- Discounts are also offered to our editors and reviewers, for more information see here.
- Special discounts can be requested by the authors of extensive review papers and monographs.
- Waivers (once per year per (co-) author for manuscripts no larger than 10 printed pages, or for the first 10 pages of a larger manuscript) are offered to:
- Retired scientists who are editors or active reviewers for this journal (1-3 reviews provided in the year before the manuscript submission).
- Scientists living and working in low-income countries (http://data.worldbank.org/income-level/low-income), if they are sole authors of a manuscript, or authors' research is funded primarily (50% or more of the work contained within the article) by an institution or organization from the eligible countries.
The journal offers also various institutional programs and membership plans to support Open Access scientific publishing. To be eligible, the author must be a corresponding author affiliated with the institution or agency.
Discounts and waivers do not accumulate.
Additional Services (Optional)
|
Optional service |
Price |
Notes |
|
Linguistic services |
€ 15 per 1800 characters |
For texts that require additional English language editing |
|
Tailored PR campaign |
€ 150* |
Press release, dedicated media and social networks promotion |
|
Paper reprints |
At cost |
On demand |
|
Auditing of the Darwin Core data associated with manuscript** |
€ 75 for datasets up to 10000 records. For large datasets (10,000 + records) please contact Dr. Bob Mesibov for pricing |
On demand |
|
Cleaning of the Darwin Core data associated with my manuscript** |
€ 225 for datasets up to 10000 records. For large datasets (10,000 + records) please contact Dr. Bob Mesibov for pricing |
On demand |
| Scientific illustrations & image processing to complement articles | ask for a quote contacting Pensoft Publishers at designer@pensoft.net | On demand |
*This service can be discounted or waived for articles of outstanding importance for the science and society.
**Pensoft reviewers do not usually have time to check through large data files included with manuscripts. If you would like us to have your data files checked, we offer the services of Pensoft editor Dr Bob Mesibov, who is also a data auditor.
Suitable data files for checking would be large tables of occurrence records or of genetic data. These can be checked for duplicate and broken records, misuse of fields, disagreements between fields, character encoding problems and incorrect or inconsistent formatting. Georeferencing can also be checked, on request. Please note that this service does not apply to taxonomic, nomenclatural or bibliographic details in data files.
Institutional and Other Membership Plans
Our plans provide additional flexibility and affordability for institutions, research groups, consortia, conference organizers and other larger research teams and organizations. Affiliated authors can publish in any Pensoft journal by using a streamlined payment interface. Pensoft’s plans are a great way to support open access publishing, while also simplifying budgeting, invoicing, and author reimbursement procedures. We offer three plans to choose from, however, if they do not quite suit your needs, we would be happy to discuss alternative arrangements with you. Please do not hesitate to contact us for a preliminary conversation about our plans!
|
Annual membership |
Pre-paid plans |
Direct billing |
|
|
Key benefits |
|
|
|
|
Additional services we can provide upon request |
|
||
Please find more details about each individual plan below. If you would like to recommend Pensoft’s plans to your institution you can fill out this simple form or contact us at info@pensoft.net and we will forward your recommendation with some additional information.
Annual Memberships
Annual memberships allow institutions to plan their publishing expenses in the beginning of the fiscal year by providing unlimited publishing in all Pensoft journals in exchange for a flat annual payment. The cost of membership depends on the total publishing output capacity of the institution and its historical publishing pattern in Pensoft journals. We will adjust the cost of your membership annually.
Pre-Paid Plans
Pre-paid plans allow institutions and / or research groups to deposit a certain amount of funds with Pensoft and make them available to affiliated researchers for covering Article Processing Charges (APCs) in any Pensoft journal. Member institutions decide whether to cover APCs in full or share the expenses with the authors. Depending on the amount members are prepared to commit, Pensoft is offering a discount on APCs per the table below. Additional funds can be added to an account at any point in time within the calendar year of purchasing the plan, while leftover funds are preserved until spent.
| Economy | Standard | Premium |
Minimum deposit | € 1,000 – 3,000 | € 3,000 – 5,000 | € 5,000 + |
Discount on APCs | 0% | 5% | 10% |
Annual awards
To recognise the research impact of our authors and the invaluable support of our editors, while also encouraging further valuable and extensive contributions to science, each January, we will be awarding the first authors1 of the three most cited 3-year-old2 articles, according to Web of Science, and the three editors3 with the most editorial tasks completed in the last calendar year.
Each awardee will receive a voucher for one free publication in the journal eligible for a submission made during the next two years.
One waiver is valid for one manuscript of standard size (up to 20 print pages), where additional pages will be charged according to our regular article processing charges. In case awarded authors do not intend to use their vouchers, they are welcome to pass the waiver to any of their co-authors.
_______________
1In case the third place is shared by multiple papers, we are awarding the most recently published one in recognition of its addressing a particularly ‘hot’ topic in the field.
2We assume that this is the minimum period for an article to accumulate a prominent amount of citations. For example, in 2023, we are awarding articles published in 2020.
3If multiple editors share the third place in the ranking, the award is given to the one with the highest all-time record at the journal.
2023 Awards
At the end of 2023, the Biodiversity Data Journal team awards the lead authors of the publications:
- Karlsson D, Hartop E, Forshage M, Jaschhof M, Ronquist F (2020) The Swedish Malaise Trap Project: A 15 Year Retrospective on a Countrywide Insect Inventory. Biodiversity Data Journal 8: e47255. https://doi.org/10.3897/BDJ.8.e47255
- Janssens SB, Couvreur TL.P, Mertens A, Dauby G, Dagallier L-PMJ, Vanden Abeele S, Vandelook F, Mascarello M, Beeckman H, Sosef M, Droissart V, van der Bank M, Maurin O, Hawthorne W, Marshall C, Réjou-Méchain M, Beina D, Baya F, Merckx V, Verstraete B, Hardy O (2020) A large-scale species level dated angiosperm phylogeny for evolutionary and ecological analyses. Biodiversity Data Journal 8: e39677. https://doi.org/10.3897/BDJ.8.e39677
- Cervellini M, Zannini P, Di Musciano M, Fattorini S, Jiménez-Alfaro B, Rocchini D, Field R, R. Vetaas O, Irl SD.H, Beierkuhnlein C, Hoffmann S, Fischer J-C, Casella L, Angelini P, Genovesi P, Nascimbene J, Chiarucci A (2020) A grid-based map for the Biogeographical Regions of Europe. Biodiversity Data Journal 8: e53720. https://doi.org/10.3897/BDJ.8.e53720
Awards from Biodiversity Data Journal also receive the three most active editors in 2023:
- Yanfeng Tong
- Alireza Zamani
- Ning Jiang
Science Communication
Our journal and the PR team at Pensoft invites authors to contribute to the communication and promotion of their published research, thereby increasing the visibility, outreach and impact of their work.
Authors are welcome to notify us whenever their institution is working on a promotional campaign about their work published in our journal. We are always happy to reshare and/or repost (where appropriate).
You can contact our PR team at dissemination@pensoft.net to discuss the communication and promotion of your research.
Tailored PR Campaign
(Paid service*)
We encourage authors, who feel that their work is of particular interest to the wider audience, to email us with a press release draft** (see template and guidelines), outlining the key findings from the study and their public impact. Then, the PR team will work with them to finalise the announcement that will be:
- Issued on the global science news service Eurekalert!
- Sent out to our media contacts from the world’s top-tier news outlets
- Posted on ARPHA’s or Pensoft’s blog
- Shared on social media via suitable ARPHA-managed accounts
Following the distribution of the press announcement, our team will be tracking the publicity across news media, blogs and social networks, in order to report back to the author(s), and reshare any prominent media content.
Request our Tailored PR campaign service by selecting it while completing your submission form and you will be contacted once your manuscript is accepted for publication. Alternatively, contact our PR team (dissemination@pensoft.net), preferably upon the acceptance of your manuscript.
* The Tailored PR campaign is an extra service charged at EUR 150. However, we would consider discounts and even full waivers for studies of particular interest for the society.
** Please note that our PR team reserves the right to edit your press release at their discretion. No press announcements will be issued until we receive the author’s final approval to do so. The service is only available for studies published within the past 3 months.
Guest Blog Post
(Free service)
Authors are strongly encouraged to promote their work and its impact on society to the audience beyond their immediate public of fellow scientists by means of storytelling in plain language. Ideally, such guest blog posts will be:
- Written from the author’s own point of view, using conversational tone;
- Written in fluent English;
- Presenting some curious background information, in order to place the discovery in context;
- Including attractive non-copyright imagery.
Request our Guest blog post service by contacting the PR department (dissemination@pensoft.net), regardless of the status of your submission, as there are no time constraints for guest blog post publication. Particularly encouraged are follow-up contributions telling the story of, for example, a research paper that has led to an important policy to be set in place; or an article that has met remarkable attention or reactions in the public sphere.
Following the necessary final touches to the guest blog post by the PR team, the contribution will be:
- Posted on ARPHA’s or Pensoft’s blog
- Shared on social media via multiple and relevant ARPHA-managed accounts
Please note that the PR team reserves the right to refuse publication of a guest blog post on the occasion that it is provided in poor English, uses considerable amount of jargon or does not abide by basic ethical standards. Our PR team reserves the right to request changes to the text related to formatting or language. No blog posts will be issued until we receive the author’s final approval to do so.
Find past guest blog posts on Pensoft’s blog here.
Video Podcast
(Free service)
To efficiently increase the outreach of their research, authors are suggested to prepare a video contribution (i.e. elevator video pitch, video abstract or topical video), where they present their work to an audience beyond their immediate public of fellow scientists by means of visual storytelling.
To do so, they are expected to send us a short (up to 02’00’’) video clip, presenting their study in a nutshell, in order to spark the viewer’s further interest in their findings and work, as well as the research topic as a whole. Ideally, such contribution will be:
- filmed in high quality, preferably with .mp4 file extension with the H.264 video codec;
- directed from the author’s own point of view, using conversational tone and minimal jargon;
- presented in fluent English or featuring English subtitles;
- accompanied by a transcript in English;
- accompanied by a short text introduction for the purposes of a blog post.
Request our Guest video contribution service by contacting the PR department (dissemination@pensoft.net), regardless of the status of your submission, since there are no time constraints for guest blog post publication.
Following the necessary final touches to the guest blog post, the contribution will be:
- Shared on Pensoft’s YouTube channel;
- Posted on ARPHA’s or Pensoft’s blog;
- Shared on social media via multiple and relevant ARPHA-managed accounts.
Please note that the PR team reserves the right to refuse distribution of a guest contribution on the occasion that it is provided in poor English, uses considerable amount of jargon or does not abide by basic ethical standards.
Custom Social Media Content
(Free service)
To help increase the visibility and outreach of their research, authors are welcome to suggest custom social media content to be distributed via suitable Pensoft- and ARPHA-managed social media accounts.
Social media posts are expected to:
- Be limited to two short sentences or 280 characters (including links);
- Be written in a conversational tone;
- Contain minimal jargon;
- Include the DOI link of the article;
- Not duplicate the title or abstract of the article;
- Include attractive non-copyright imagery;
- Possibly include up to 10 social media accounts, e.g. co-authors (Twitter only), affiliations, funding bodies etc. relevant to the study.
Request our Custom social media content service by contacting our PR department (dissemination@pensoft.net).
Please note that our PR team reserves the right to edit your text at their discretion.
Media Center
Follow Biodiversity Data Journal on Twitter and Facebook.
Learn about some of the most notable research published in Biodiversity Data Journal on Pensoft's blog.
See top news stories from around the globe, mentioning research published in Biodiversity Data Journal on Fox News, USA Today, CNET, New York Post, РИА Новости (RIA Novosti), Detroit Free Press, Der Standard, Gazeta.ru, International Business Times and The Telegraph (India).
Boost the reach of your paper(s) to a larger audience by making the most of Pensoft's science communication services.
Download journal leaflet.
Download journal logo.
Web Services
OAI-PMH - oai_dc: https://bdj.pensoft.net/oai.php?verb=ListRecords&set=bdj&metadataPrefix=oai_dc
OAI-PMH - mods: https://bdj.pensoft.net/oai.php?verb=ListRecords&set=bdj&metadataPrefix=mods
RSS for metadata: https://bdj.pensoft.net/rss.php
Journal Info
| Journal Name | Biodiversity Data Journal |
|---|---|
| Journal URL | https://bdj.pensoft.net/ |
| ISSN (online) | 1314-2828 |
| ISSN (print) | 1314-2836 |
| Content Provider | ARPHA |
| Publisher | Pensoft Publishers |
| Journal Owner | Pensoft Publishers |
| Owner URL | https://pensoft.net |
| Start Year | 2013 |
| Review Type | single-blind |
| Publication Frequency | continuous |
| APC | Accepted manuscripts are subject to APC (for more details see here) |
| License | Creative Commons Attribution License (CC BY 4.0) |